 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | kfl00028_0250 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Klebsormidiophyceae; Klebsormidiales; Klebsormidiaceae; Klebsormidium
|
||||||||
| Family | GATA | ||||||||
| Protein Properties | Length: 416aa MW: 44816.5 Da PI: 5.7657 | ||||||||
| Description | GATA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-B_box | 22.7 | 2e-07 | 14 | 54 | 4 | 38 |
zf-B_box 4 rkCpeHeekelqlfCedCqqllCedClleeHkg......Ht 38
kC+ ++e +++lfC +++++C++C ++ H g H+
kfl00028_0250 14 LKCDVCQEARAILFCPRDRSCFCQQCDFAAHAGaqstacHE 54
68*****************************8777777776 PP
| |||||||
| 2 | GATA | 52 | 9.5e-17 | 312 | 346 | 1 | 35 |
GATA 1 CsnCgttkTplWRrgpdgnktLCnaCGlyyrkkgl 35
C +C +tkTp+WR gp g ktLCnaCG++y++ +l
kfl00028_0250 312 CLHCASTKTPQWRAGPMGVKTLCNACGVRYKSGRL 346
99*****************************9885 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50119 | 8.887 | 11 | 58 | IPR000315 | B-box-type zinc finger |
| SMART | SM00336 | 0.0041 | 11 | 58 | IPR000315 | B-box-type zinc finger |
| CDD | cd00021 | 1.55E-6 | 15 | 46 | No hit | No description |
| PROSITE profile | PS50114 | 11.807 | 306 | 342 | IPR000679 | Zinc finger, GATA-type |
| SMART | SM00401 | 1.0E-14 | 306 | 356 | IPR000679 | Zinc finger, GATA-type |
| SuperFamily | SSF57716 | 5.23E-12 | 308 | 344 | No hit | No description |
| Gene3D | G3DSA:3.30.50.10 | 4.0E-14 | 310 | 343 | IPR013088 | Zinc finger, NHR/GATA-type |
| CDD | cd00202 | 8.07E-12 | 311 | 366 | No hit | No description |
| PROSITE pattern | PS00344 | 0 | 312 | 337 | IPR000679 | Zinc finger, GATA-type |
| Pfam | PF00320 | 1.4E-14 | 312 | 346 | IPR000679 | Zinc finger, GATA-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009416 | Biological Process | response to light stimulus | ||||
| GO:0005622 | Cellular Component | intracellular | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 416 aa Download sequence Send to blast |
MLPGSAALPA TDYLKCDVCQ EARAILFCPR DRSCFCQQCD FAAHAGAQST ACHERYALAQ 60 TKITRMPLEK PASMPVAASL PQRVPPGCTP SDEEAFLVDD LLDLSEAKGE SRVDSAFSPS 120 QEQQGQVEQQ REPEREPPSA SGATSWDSEE VGEVGDLCDL PCDDLAELDW LSTFVENSYS 180 SPDGVPLSAP DGGGGLAQGA EHPLKGVGMR SKRQRANPSR TWSAELLIPR DDDLPKRPKF 240 NRSWSTSNLA RPSKPFHRSA IPEAASCSDL SALERACTQS QVGIECEPAP PEPPAGIAGA 300 NFEQLPHPLR RCLHCASTKT PQWRAGPMGV KTLCNACGVR YKSGRLVADY RPAGSPGFAE 360 GNGNKPNYGG KGAPPCPPIA RKKLLIPKAA RRMVPKPKRE FGVEELSGYS CDEMEM |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00048 | PBM | Transfer from AT4G32890 | Download |
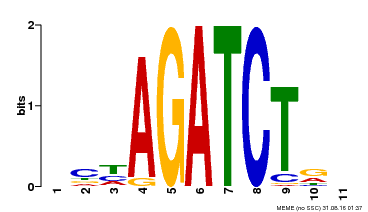
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| TrEMBL | A0A1Y1HL38 | 0.0 | A0A1Y1HL38_KLENI; Uncharacterized protein | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP68 | 17 | 287 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G32890.1 | 8e-28 | GATA transcription factor 9 | ||||



