 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | WALNUT_00026408-RA | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fagales; Juglandaceae; Juglans
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 1003aa MW: 111627 Da PI: 5.7962 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 125.8 | 1.8e-39 | 154 | 231 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+Cqv++C adls+ak+yhrrh+vCe+hska+++lv + +qrfCqqCsrf el+efDe+krsCrrrLa+hn+rrrk+++
WALNUT_00026408-RA 154 ICQVDHCGADLSHAKDYHRRHRVCEMHSKATKALVGNSMQRFCQQCSRFQELQEFDEGKRSCRRRLAGHNKRRRKTNP 231
5**************************************************************************976 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 7.0E-32 | 148 | 216 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 31.094 | 152 | 229 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 2.35E-36 | 153 | 235 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 6.7E-29 | 155 | 228 | IPR004333 | Transcription factor, SBP-box |
| Gene3D | G3DSA:1.25.40.20 | 2.8E-6 | 749 | 905 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 8.7E-7 | 780 | 889 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 2.27E-6 | 781 | 889 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1003 aa Download sequence Send to blast |
MEAGFGGEAH HFYAMSAADL RVMGKRSLEW DLNEWKWDGD LFMASQLNPV PSGCMDQQFL 60 PLGSGIPLTG GSSNSSSSCS DEVNLEVERG KRELEKRRRV IVLEDDNSND EEEGTLALKL 120 GGQDYQVAER EARNWEGMDG KKTKLVGGTS NRAICQVDHC GADLSHAKDY HRRHRVCEMH 180 SKATKALVGN SMQRFCQQCS RFQELQEFDE GKRSCRRRLA GHNKRRRKTN PDTVVNGSTM 240 NDDQTSGYIL ISLLKILSNM HSNRSDQATD QDLLSHFLRS LASQTGEQVG KNISGILKEA 300 PNSQNAGNSL GNSELVSTLF TNGPHDPSRT FKDHHPVPVT EMPQQGLRVD EARGGDIQTA 360 SSSKPSILSS PPVYSEVRDS TARQIKMNNF DLNDIYIDSD DGMEDIDRLP DPANMGASSL 420 DSHQSSPPQT SGHSDSVSAQ SPTSSSGDAQ SRTDRIVFKL FGKEPNDFPL VLRAQILDWL 480 SHSPTDIESY IRPGCIILTI YLCQSEAAWE ELCHDLSSSL NRLLDVSDDS FWRTGWVYIR 540 VQHQIAFIYN GQVVIDTSLP IRSNNYGQIL SVKPIAISTS ERAQFSVKGI NLTRPATRLL 600 CALEGNCQVE ENTHDSTDAL DNFEDHDELQ SINFSCSIHA VNGRGFIEIE DHGFSSSFFP 660 FIVAEEDVCS EIRALESALE STETSADFVG TRKMEAKNQA LDFIHEMGWL LHRSQLKSRL 720 GRMDPNEDLF PLKRFKWLME FSMDQDWCAV VRKLLDILLD GTVGAGEHPS MNLALAEMGL 780 LHRAVRRNSR ALVELLLRYQ VSNELRSESV ELFGGGNESF LFRPDVLGPA GLTPLHIAAG 840 KDGSENVLDA LTNDLKMVGI EAWKSARDST GSTPEDYARL RGHFSYIHLV QRKINKRSAA 900 GHVVLDIPSA PSDCNANQKQ SNELTTSFEI GAESRRVRQH CKLCDQKLAY GTACRSLVYR 960 PVMLSLVTIA VVCVCVALLF KSSPEVLYVF RPFRWEMLGY GTS |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 3e-30 | 148 | 228 | 4 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 95 | 99 | KRRRV |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3'. {ECO:0000269|PubMed:16554053}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00633 | PBM | Transfer from PK06791.1 | Download |
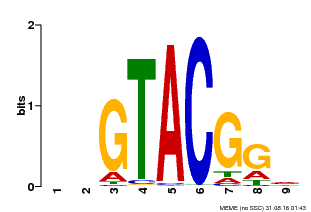
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018832330.1 | 0.0 | PREDICTED: squamosa promoter-binding-like protein 1 isoform X2 | ||||
| Swissprot | Q9S7P5 | 0.0 | SPL12_ARATH; Squamosa promoter-binding-like protein 12 | ||||
| TrEMBL | A0A2I4FL14 | 0.0 | A0A2I4FL14_JUGRE; squamosa promoter-binding-like protein 1 isoform X2 | ||||
| STRING | POPTR_0002s18970.1 | 0.0 | (Populus trichocarpa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1539 | 34 | 95 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47070.1 | 0.0 | squamosa promoter binding protein-like 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



