 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | WALNUT_00022582-RA | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fagales; Juglandaceae; Juglans
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 701aa MW: 77026.5 Da PI: 6.7014 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 74.3 | 1.4e-23 | 121 | 222 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT.......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkl 84
f k+lt+sd+++ g +++p+ +ae++ ++ + t+ +d +g++W++++iyr++++r++lt+GW+ Fv+ ++L +gD++vF
WALNUT_00022582-RA 121 FAKTLTQSDANNGGGFSVPRYCAETIfprldysADPPVQ--TILAKDVHGETWKFRHIYRGTPRRHLLTTGWSTFVNHKKLVAGDSIVFL- 208
89****************************996555555..89***********************************************. PP
-SSSEE..EEEEE-S CS
B3 85 dgrsefelvvkvfrk 99
+ +++ l+v+++r+
WALNUT_00022582-RA 209 -RAENGDLCVGIRRA 222
.4488999****997 PP
| |||||||
| 2 | Auxin_resp | 113.9 | 1.5e-37 | 292 | 375 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrl.sGtvvgvsdldpvrWpnSkWrsLk 83
aa+ a+++++FevvY+Prast+eF+vk++ v++al++++++GmRfkmafetedss++++ +Gt+++v+ +dp+rWp+S+Wr+L+
WALNUT_00022582-RA 292 AATLAANGKAFEVVYYPRASTPEFCVKASMVKAALQIRWCAGMRFKMAFETEDSSRISWfMGTISSVQVADPLRWPESPWRLLQ 375
68899*****************************************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.40.330.10 | 1.0E-37 | 112 | 225 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 4.71E-29 | 115 | 224 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 2.96E-21 | 120 | 221 | No hit | No description |
| PROSITE profile | PS50863 | 13.112 | 121 | 223 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 1.7E-21 | 121 | 223 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 8.8E-22 | 121 | 222 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 4.8E-33 | 292 | 375 | IPR010525 | Auxin response factor |
| PROSITE profile | PS51745 | 21.645 | 615 | 698 | IPR000270 | PB1 domain |
| SuperFamily | SSF54277 | 2.16E-5 | 618 | 689 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007389 | Biological Process | pattern specification process | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0048829 | Biological Process | root cap development | ||||
| GO:0051301 | Biological Process | cell division | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 701 aa Download sequence Send to blast |
MITFMDSKEK LKEVEKCLDP QLWHACAGGM VQMPSVNAKV FYFPQGHAEH ACGNVDFRSC 60 PRVVPFIPCR VCAVKYMADP ETDEVFTKMR LVPVNANELD FEDDGIGGIN GSEAQEKPAS 120 FAKTLTQSDA NNGGGFSVPR YCAETIFPRL DYSADPPVQT ILAKDVHGET WKFRHIYRGT 180 PRRHLLTTGW STFVNHKKLV AGDSIVFLRA ENGDLCVGIR RAKRGIGGGP EMTSGWNPAG 240 GSCAMPYGGF SAFLRDGESK LMRNCNSNGN GSSSNGNLMV RGKVRPESVI EAATLAANGK 300 AFEVVYYPRA STPEFCVKAS MVKAALQIRW CAGMRFKMAF ETEDSSRISW FMGTISSVQV 360 ADPLRWPESP WRLLQVTWDE PDLLQNVKCV SPWLVELVSN IPAMHLSPFS PPRKKLRLPQ 420 LPDFPLDGQL HMPTFSGNLL GPSSPFSCLP ENIPAGMQGA RHAHYGLSLS DLHLNKLQSG 480 LFPAGFPRLD HTATPARAPN NSIFQKPSMS ENVSCFLTMA HSQTPKKPDD VKSPQFVLFG 540 QPILTEQQIS LSCSGDTVSP VGTGNSSSDG NADKVTNFSD GSGCAHNRGI VERSSCEGLQ 600 WYKDNCQETE PNLETGHCKV FMESEDVGRS LDLSLLGSYE ELYRKLANMF SIENSETLSQ 660 VLYRDTTGAV KLMGDEPFSD FMKTARRLTI LMDSSSDNVG M |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldu_A | 4e-77 | 18 | 396 | 51 | 388 | Auxin response factor 5 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00461 | DAP | Transfer from AT4G30080 | Download |
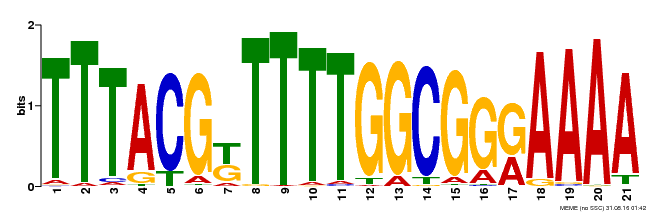
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018838364.1 | 0.0 | PREDICTED: auxin response factor 18-like | ||||
| Refseq | XP_018838365.1 | 0.0 | PREDICTED: auxin response factor 18-like | ||||
| Swissprot | Q653H7 | 0.0 | ARFR_ORYSJ; Auxin response factor 18 | ||||
| TrEMBL | A0A2I4G3C5 | 0.0 | A0A2I4G3C5_JUGRE; Auxin response factor | ||||
| STRING | EOY07838 | 0.0 | (Theobroma cacao) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF637 | 34 | 138 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G30080.1 | 0.0 | auxin response factor 16 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



