 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | WALNUT_00021643-RA | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fagales; Juglandaceae; Juglans
|
||||||||
| Family | GATA | ||||||||
| Protein Properties | Length: 319aa MW: 34255.1 Da PI: 4.6249 | ||||||||
| Description | GATA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GATA | 50.2 | 3.5e-16 | 229 | 265 | 1 | 35 |
GATA 1 CsnCgttk..TplWRrgpdgnktLCnaCGlyyrkkgl 35
C++Cg ++ Tp++Rrgp g++tLCnaCGl+++ kg+
WALNUT_00021643-RA 229 CTHCGISSksTPMMRRGPTGPRTLCNACGLKWANKGV 265
*****99999************************997 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00979 | 4.1E-4 | 78 | 110 | IPR010399 | Tify domain |
| PROSITE profile | PS51320 | 10.049 | 78 | 113 | IPR010399 | Tify domain |
| Pfam | PF06200 | 3.5E-8 | 82 | 104 | IPR010399 | Tify domain |
| Pfam | PF06203 | 2.5E-16 | 156 | 198 | IPR010402 | CCT domain |
| PROSITE profile | PS51017 | 13.314 | 156 | 198 | IPR010402 | CCT domain |
| SMART | SM00401 | 2.0E-12 | 223 | 276 | IPR000679 | Zinc finger, GATA-type |
| PROSITE profile | PS50114 | 9.433 | 223 | 280 | IPR000679 | Zinc finger, GATA-type |
| SuperFamily | SSF57716 | 6.65E-12 | 226 | 274 | No hit | No description |
| Gene3D | G3DSA:3.30.50.10 | 1.4E-15 | 227 | 270 | IPR013088 | Zinc finger, NHR/GATA-type |
| CDD | cd00202 | 4.99E-15 | 228 | 271 | No hit | No description |
| Pfam | PF00320 | 6.2E-14 | 229 | 265 | IPR000679 | Zinc finger, GATA-type |
| PROSITE pattern | PS00344 | 0 | 229 | 256 | IPR000679 | Zinc finger, GATA-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 319 aa Download sequence Send to blast |
MPESNRQASM FNSGAMTNNI ESQIEDEDED VAGGEESIDN PQIRFEDGAV AVTFNGGVQE 60 VAPNTIYVPS SDYPPVAGNG GADQLTLSFQ GEVYVFDAVS PDKVLVFAYG WLLNFVNLLL 120 LLGGYEIPSG IPAMGTVPLN QRGMNDFPGR SIQPQRAASL SRFREKRKER CFDKKIRYSV 180 RKEVALRMQR KKGQFTSSKA ISDEAGSDSS VYGATQGPGQ DDTMQETSCT HCGISSKSTP 240 MMRRGPTGPR TLCNACGLKW ANKGVLRDLS KVSTMGVQDP SLKATEQGDG DASESDAVTP 300 VAEIVSTANG DNSAVTAER |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that specifically binds 5'-GATA-3' or 5'-GAT-3' motifs within gene promoters. {ECO:0000250, ECO:0000269|PubMed:14966217}. | |||||
| UniProt | Transcriptional activator that specifically binds 5'-GATA-3' or 5'-GAT-3' motifs within gene promoters. {ECO:0000250, ECO:0000269|PubMed:14966217}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00198 | DAP | Transfer from AT1G51600 | Download |
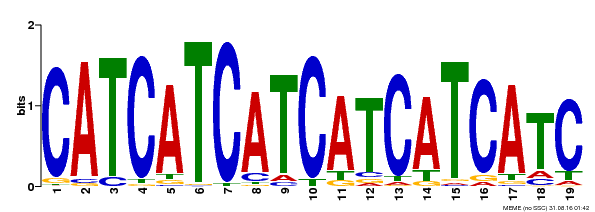
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018811236.1 | 0.0 | PREDICTED: GATA transcription factor 24-like | ||||
| Swissprot | Q8GXL7 | 4e-71 | GAT24_ARATH; GATA transcription factor 24 | ||||
| Swissprot | Q8H1G0 | 6e-71 | GAT28_ARATH; GATA transcription factor 28 | ||||
| TrEMBL | A0A2I4DVR1 | 0.0 | A0A2I4DVR1_JUGRE; GATA transcription factor 24-like | ||||
| STRING | XP_010112233.1 | 1e-151 | (Morus notabilis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF2836 | 33 | 68 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G21175.2 | 3e-71 | ZIM-like 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



