 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | WALNUT_00020183-RA | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fagales; Juglandaceae; Juglans
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 251aa MW: 28670.3 Da PI: 6.3037 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 56.4 | 5.1e-18 | 48 | 101 | 3 | 56 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k+++++ eq+++Le+ Fe +++ e++ +LA+ lgL+ rq+ +WFqNrRa++k
WALNUT_00020183-RA 48 KKRRLNMEQVKTLEKNFELGNKLEPERKMQLARALGLQPRQIAIWFQNRRARWK 101
456899***********************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 128.8 | 2.2e-41 | 47 | 137 | 1 | 91 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLreel 91
ekkrrl+ eqvk+LE++Fe +kLeperK++lar+Lglqprq+a+WFqnrRAR+ktkqlEkdy+ Lkr+++a+k++n+ L++++++L++e+
WALNUT_00020183-RA 47 EKKRRLNMEQVKTLEKNFELGNKLEPERKMQLARALGLQPRQIAIWFQNRRARWKTKQLEKDYDLLKRQFEAIKADNDVLQAQNQKLQAEI 137
69**************************************************************************************987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 3.4E-19 | 24 | 104 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 1.5E-19 | 39 | 105 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.119 | 43 | 103 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.8E-17 | 46 | 107 | IPR001356 | Homeobox domain |
| Pfam | PF00046 | 2.3E-15 | 48 | 101 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.64E-15 | 48 | 104 | No hit | No description |
| PRINTS | PR00031 | 9.1E-6 | 74 | 83 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 78 | 101 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 9.1E-6 | 83 | 99 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 4.4E-15 | 103 | 143 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009744 | Biological Process | response to sucrose | ||||
| GO:0048826 | Biological Process | cotyledon morphogenesis | ||||
| GO:0080022 | Biological Process | primary root development | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 251 aa Download sequence Send to blast |
MGFKCFFVDS GVASFLGKRS LSFSGVELGE EGNGEDDLSD DGSQVGEKKR RLNMEQVKTL 60 EKNFELGNKL EPERKMQLAR ALGLQPRQIA IWFQNRRARW KTKQLEKDYD LLKRQFEAIK 120 ADNDVLQAQN QKLQAEILAL KNREPTESIN LNKETEGSCS NRSENSSEIK LDISRTPAID 180 SPLSTHPISR TLFPSSVRPP GVAQLFQNSS RPDLQCQKMD QMVKEESLSN MFCGMDDQSA 240 FWPWLEQHHF N |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 95 | 103 | RRARWKTKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that may act in the sucrose-signaling pathway. {ECO:0000269|PubMed:11292072}. | |||||
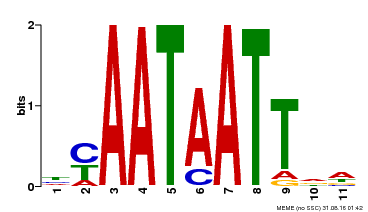
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00225 | DAP | Transfer from AT1G69780 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018857022.1 | 1e-177 | PREDICTED: homeobox-leucine zipper protein ATHB-13-like | ||||
| Swissprot | Q8LC03 | 1e-106 | ATB13_ARATH; Homeobox-leucine zipper protein ATHB-13 | ||||
| TrEMBL | A0A2I4HLP7 | 1e-176 | A0A2I4HLP7_JUGRE; homeobox-leucine zipper protein ATHB-13-like | ||||
| STRING | VIT_01s0026g01950.t01 | 1e-155 | (Vitis vinifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1271 | 33 | 106 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G69780.1 | 9e-99 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



