 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | WALNUT_00019810-RA | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fagales; Juglandaceae; Juglans
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 623aa MW: 70181.6 Da PI: 8.0941 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 172 | 1.8e-53 | 308 | 437 | 1 | 128 |
NAM 1 lppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLpk..kvka.eekewyfFskrdkkyatgkrknratksgyWkatgk 88
+ppGfrFhPtdeel+ +yL+kkv+ + ++l +vi+evd++k+ePwdL++ ++ + ++ewyfFs++dkky+tg+r+nrat++g+Wkatg+
WALNUT_00019810-RA 308 VPPGFRFHPTDEELLYYYLRKKVSYEAIDL-DVIREVDLNKLEPWDLKEkcRIGSgPQNEWYFFSHKDKKYPTGTRTNRATTAGFWKATGR 397
69****************************.9***************952444443456******************************** PP
NAM 89 dkevlskkgelvglkktLvfykgrapkgektdWvmheyrl 128
dk+++ ++ + +g++ktLvfy+grap+g+ktdW+mheyrl
WALNUT_00019810-RA 398 DKAIHLSNAKSIGMRKTLVFYTGRAPHGQKTDWIMHEYRL 437
**************************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF53335 | 1.11E-8 | 105 | 268 | IPR029063 | S-adenosyl-L-methionine-dependent methyltransferase |
| Gene3D | G3DSA:3.40.50.150 | 2.2E-9 | 173 | 256 | IPR029063 | S-adenosyl-L-methionine-dependent methyltransferase |
| SuperFamily | SSF101941 | 3.27E-59 | 299 | 457 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 58.144 | 308 | 457 | IPR003441 | NAC domain |
| Pfam | PF02365 | 1.1E-27 | 309 | 437 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003002 | Biological Process | regionalization | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009834 | Biological Process | plant-type secondary cell wall biogenesis | ||||
| GO:0010455 | Biological Process | positive regulation of cell fate commitment | ||||
| GO:0048829 | Biological Process | root cap development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 623 aa Download sequence Send to blast |
MHAYMATYQR THVAGCVLEF TKLGLHHVIS DETEKGKRKA KGKGSFAFRT NCSISISKER 60 WSSSSRSRSC VGNLKVSAAK SQEPVDSSRA EATQEEEEGD DENYRVLSAV KSSFNDILIL 120 DSSKSRILLL DSTHNVHSIL YKDQKWTGSY WDEFASLPAI VPKGPIAIFG LLIDKAREYL 180 GLSYLEKHTV AGGILNVHIG DALSQSVNVS EGYAGIVIDL FSEGKVLPQL QEVNTWLELT 240 DRLMPKGRLM VNCGGINELS WKRMPKRDGE NYLALTGPLP DLASWSAMVP GLLIKDQKMM 300 PGNGQLSVPP GFRFHPTDEE LLYYYLRKKV SYEAIDLDVI REVDLNKLEP WDLKEKCRIG 360 SGPQNEWYFF SHKDKKYPTG TRTNRATTAG FWKATGRDKA IHLSNAKSIG MRKTLVFYTG 420 RAPHGQKTDW IMHEYRLDDD NSQVQEDGWV VCRVFKKKNH CRVFQPESAQ EAEHFTQMKA 480 SGSAPVLLEP KQTHLRPLYD YKFDGPMHLP QLFSPESAFA QPSFISPASL NTMDIQCSQN 540 LLRLTAGGCA GLMHQERFNG DWSFLDKLLA SHHQSMDHHQ HPQSKCNPSS HQVPDHVGSS 600 SSTSQKFPFQ YLGCETDILK FSK |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 3ulx_A | 2e-49 | 304 | 459 | 11 | 170 | Stress-induced transcription factor NAC1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription regulator. Together with BRN1 and BRN2, regulates cellular maturation of root cap. Represses stem cell-like divisions in the root cap daughter cells, and thus promotes daughter cell fate. Inhibits expression of its positive regulator FEZ in a feedback loop for controlled switches in cell division plane. Promotes the expression of genes involved in secondary cell walls (SCW) biosynthesis. {ECO:0000269|PubMed:19081078, ECO:0000269|PubMed:20197506}. | |||||
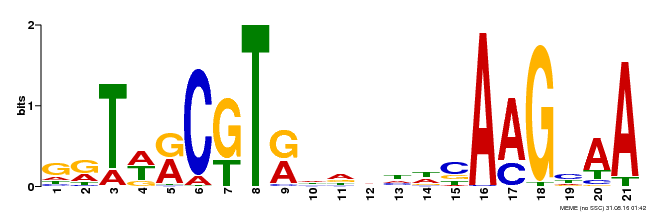
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00250 | DAP | Transfer from AT1G79580 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By FEZ in oriented-divised root cap stem cells. {ECO:0000269|PubMed:19081078}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018811818.1 | 0.0 | PREDICTED: protein SOMBRERO-like isoform X1 | ||||
| Swissprot | Q9MA17 | 1e-126 | SMB_ARATH; Protein SOMBRERO | ||||
| TrEMBL | A0A2I4DXD7 | 0.0 | A0A2I4DXD7_JUGRE; protein SOMBRERO-like isoform X1 | ||||
| STRING | VIT_19s0085g01210.t01 | 0.0 | (Vitis vinifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF7204 | 34 | 47 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G79580.3 | 1e-118 | NAC family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



