 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | WALNUT_00019069-RA | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fagales; Juglandaceae; Juglans
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1105aa MW: 123601 Da PI: 5.3916 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 168.7 | 9.1e-53 | 30 | 132 | 3 | 105 |
CG-1 3 kekkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenpt 93
+ ++rwl++ ei++iL n++k+++++e+++rp+sgsl+L++rk++ryfrkDG++w+kkkdgktv+E+hekLKvg+v+vl+cyYah+een+t
WALNUT_00019069-RA 30 EARSRWLRPAEICEILRNYQKFHIASEPPNRPPSGSLFLFDRKVLRYFRKDGHNWRKKKDGKTVKEAHEKLKVGSVDVLHCYYAHGEENET 120
459**************************************************************************************** PP
CG-1 94 fqrrcywlLeee 105
fqrr+yw+Le++
WALNUT_00019069-RA 121 FQRRSYWMLEQN 132
**********98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 72.576 | 24 | 150 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 2.2E-69 | 27 | 407 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 6.1E-46 | 30 | 133 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 8.7E-8 | 486 | 591 | IPR013783 | Immunoglobulin-like fold |
| Pfam | PF01833 | 8.5E-7 | 505 | 578 | IPR002909 | IPT domain |
| SuperFamily | SSF81296 | 4.36E-18 | 505 | 591 | IPR014756 | Immunoglobulin E-set |
| SuperFamily | SSF48403 | 5.91E-14 | 689 | 808 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 13.575 | 689 | 800 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 1.4E-13 | 691 | 810 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 2.25E-11 | 695 | 803 | No hit | No description |
| SuperFamily | SSF52540 | 1.06E-8 | 920 | 974 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.056 | 923 | 945 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0015 | 925 | 944 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 7.913 | 925 | 953 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0076 | 946 | 968 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.395 | 947 | 971 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 4.9E-4 | 949 | 968 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1105 aa Download sequence Send to blast |
MPISRVCFMG LDSVLLGSHL FLDIQQILAE ARSRWLRPAE ICEILRNYQK FHIASEPPNR 60 PPSGSLFLFD RKVLRYFRKD GHNWRKKKDG KTVKEAHEKL KVGSVDVLHC YYAHGEENET 120 FQRRSYWMLE QNRDLYSSLP KRLLFGFFMG NRTNIGGIRE RDEVASDFQK DSPLSSSFPT 180 NHKGAPSGST DSPSPTNSVT SLCEDADSED SYQASSRSRF LESPSMENGP LTDRMDAGGA 240 SSYFVHPSAD VQKDGPRDSD GVTHATGSED AIDLASWEEV LENSRGFHTS VTSRASSEPQ 300 STLTDIFLGQ ENMTIGDNLD GESVLKEEFQ NTVPRQPNWQ ISFGDNSVHL STWPMNQLSN 360 FELANDLGTA LFEPGTLDMN AISAPEPFST HPVQQNEQRV QHHLQTQLTN AESQFSIKSN 420 SQTDMLFEGT NGNYALTLRK PLLDGEEALK KVDSFSRWVT RELGDVDDLH LQSSSGLSWS 480 TVECGNVVDD SSLSPSLSQD QLFSILDFSP KWAYTDSETE VLITGTFLKS LPEVTKCNWS 540 CMFGEVEVPA EVLANGILCC HTPPHTVGQV PFYITCSNRV ACSEVREFDY RVGSIKDIDI 600 AVIYSGAANE MDLHLRLERL LSLSSVVPPS HVVESVTVKQ ELISKIITLK EEEEYNQTLE 660 PILQRDLSQH EVKEYLLKLL LKEKLYSWLL RKVCEDGKGP SILDDDGQGV IHLAAALGYD 720 WAIKPIVTAE VSINFRDING WTALHWAASC GRQDPLNLIL VEQTVAFLVS LGAAPGALTD 780 PSPEFPLGKT PADLASGKGH KGISGFIAES SLTSYLKSLT MNDQKEDGMP EVSGTKGVLT 840 ISERTATPVN YVDAADALSL KDSLNAVCNA TQAADRIHQM FRMHSFERRQ ITQHDDGGGS 900 LSDECALSLV SAKTRKPGQS DRLVHAAATH IQKKYRGWKK RKEFLTIRQR IVKIQAHVRG 960 HQVRKQYRTI VWSVGILEKV ILRWRRKGIG LRGFRPDVHK DPKPQVVPST EDDYDFLKEG 1020 RKQTEERLQK ALTRVKSMVQ YPEGRAQYRR LLTVVEGFRE TKAGHMVLNS PEETGDGDED 1080 EIDINALLDD EMLMDDEAFM SIAFE |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3' (By similarity). Regulates transcriptional activity in response to calcium signals (Probable). Binds calmodulin in a calcium-dependent manner (By similarity). Involved in freezing tolerance in association with CAMTA1 and CAMTA3. Contributes together with CAMTA1 and CAMTA3 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:23581962). Involved together with CAMTA3 and CAMTA4 in the positive regulation of a general stress response (PubMed:25039701). Involved in tolerance to aluminum. Binds to the promoter of ALMT1 transporter and contributes to the positive regulation of aluminum-induced expression of ALMT1 (PubMed:25627216). {ECO:0000250|UniProtKB:Q8GSA7, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25627216, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
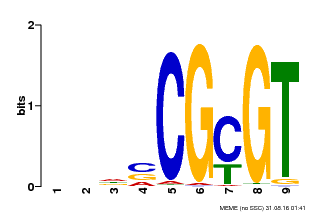
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salt, wounding, abscisic acid, H(2)O(2) and salicylic acid (PubMed:12218065). Induced by aluminum (PubMed:25627216). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:25627216}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018836863.1 | 0.0 | PREDICTED: calmodulin-binding transcription activator 2-like isoform X2 | ||||
| Swissprot | Q6NPP4 | 0.0 | CMTA2_ARATH; Calmodulin-binding transcription activator 2 | ||||
| TrEMBL | A0A2I4FYZ6 | 0.0 | A0A2I4FYZ6_JUGRE; calmodulin-binding transcription activator 2-like isoform X2 | ||||
| STRING | XP_008237378.1 | 0.0 | (Prunus mume) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF7406 | 26 | 42 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G64220.2 | 0.0 | Calmodulin-binding transcription activator protein with CG-1 and Ankyrin domains | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



