 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | WALNUT_00017585-RA | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fagales; Juglandaceae; Juglans
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 308aa MW: 34959.6 Da PI: 7.27 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 32.6 | 2.3e-10 | 22 | 49 | 3 | 30 |
NAM 3 pGfrFhPtdeelvveyLkkkvegkklel 30
pGfrFhPtdeelv +yL++kve+k++++
WALNUT_00017585-RA 22 PGFRFHPTDEELVGFYLRRKVEKKPIQA 49
9***********************9887 PP
| |||||||
| 2 | NAM | 87.9 | 1.9e-27 | 54 | 137 | 53 | 128 |
NAM 53 aeekewyfFskrdkkyatgkrknratksgyWkatgkdkevlskkgel........vglkktLvfykgrapkgektdWvmheyrl 128
a+++e yfF+kr +ky+++ r+nr+t sg+Wkatg dk+v+s+kg++ +glkktLv+y+g+a kg+ktdW+mhe+rl
WALNUT_00017585-RA 54 AKDTEEYFFCKRGRKYRNSIRPNRVTGSGFWKATGIDKQVYSAKGSAdrhshhvcIGLKKTLVYYRGSAGKGTKTDWMMHEFRL 137
567899************************************9444455566777***************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 2.48E-41 | 20 | 167 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 31.137 | 20 | 168 | IPR003441 | NAC domain |
| Pfam | PF02365 | 8.3E-25 | 22 | 137 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 308 aa Download sequence Send to blast |
MSSTMQLDTS TEDDGHDEVP LPGFRFHPTD EELVGFYLRR KVEKKPIQAA GSTAKDTEEY 60 FFCKRGRKYR NSIRPNRVTG SGFWKATGID KQVYSAKGSA DRHSHHVCIG LKKTLVYYRG 120 SAGKGTKTDW MMHEFRLPAD HDTNTNLSNS KSTAPEAEIW TLCRIFERNV SHRKYTPDWR 180 ALSAKRHTAN SANYSSSEAC SLESNNREGY INFESSTTTP FVYYNDHDHN ERKPVVNHLH 240 VDEIRNQLHV GQLSTTAPVA HERPPSSSGF PIGSPDGPDE NYHALFDCGN WDDELRSVVE 300 FALNPSRM |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ut4_A | 4e-30 | 22 | 174 | 19 | 171 | NO APICAL MERISTEM PROTEIN |
| 1ut4_B | 4e-30 | 22 | 174 | 19 | 171 | NO APICAL MERISTEM PROTEIN |
| 1ut7_A | 4e-30 | 22 | 174 | 19 | 171 | NO APICAL MERISTEM PROTEIN |
| 1ut7_B | 4e-30 | 22 | 174 | 19 | 171 | NO APICAL MERISTEM PROTEIN |
| 3swm_A | 4e-30 | 22 | 174 | 22 | 174 | NAC domain-containing protein 19 |
| 3swm_B | 4e-30 | 22 | 174 | 22 | 174 | NAC domain-containing protein 19 |
| 3swm_C | 4e-30 | 22 | 174 | 22 | 174 | NAC domain-containing protein 19 |
| 3swm_D | 4e-30 | 22 | 174 | 22 | 174 | NAC domain-containing protein 19 |
| 3swp_A | 4e-30 | 22 | 174 | 22 | 174 | NAC domain-containing protein 19 |
| 3swp_B | 4e-30 | 22 | 174 | 22 | 174 | NAC domain-containing protein 19 |
| 3swp_C | 4e-30 | 22 | 174 | 22 | 174 | NAC domain-containing protein 19 |
| 3swp_D | 4e-30 | 22 | 174 | 22 | 174 | NAC domain-containing protein 19 |
| 4dul_A | 4e-30 | 22 | 174 | 19 | 171 | NAC domain-containing protein 19 |
| 4dul_B | 4e-30 | 22 | 174 | 19 | 171 | NAC domain-containing protein 19 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
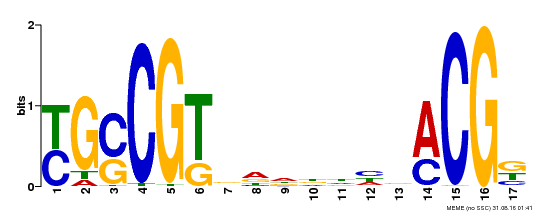
| UniProt | Transcription factor that binds to the 5'- RRYGCCGT-3' consensus core sequence. Central longevity regulator. Negative regulator of leaf senescence. Modulates cellular H(2)O(2) levels and enhances tolerance to various abiotic stresses through the regulation of DREB2A. {ECO:0000269|PubMed:22345491}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00353 | DAP | Transfer from AT3G12910 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated by H(2)O(2), paraquat, ozone, 3-aminotriazole and salt stress. {ECO:0000269|PubMed:22345491}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | JQ618146 | 1e-163 | JQ618146.1 Juglans mandshurica clone NS21 microsatellite sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018815581.1 | 0.0 | PREDICTED: transcription factor JUNGBRUNNEN 1-like, partial | ||||
| Swissprot | Q9SK55 | 2e-62 | NAC42_ARATH; Transcription factor JUNGBRUNNEN 1 | ||||
| TrEMBL | A0A2I4E856 | 0.0 | A0A2I4E856_JUGRE; transcription factor JUNGBRUNNEN 1-like | ||||
| STRING | cassava4.1_033023m | 1e-102 | (Manihot esculenta) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1013 | 34 | 112 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G12910.1 | 3e-77 | NAC family protein | ||||



