 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | WALNUT_00015337-RA | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fagales; Juglandaceae; Juglans
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 279aa MW: 30709.5 Da PI: 4.7246 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 64 | 3e-20 | 119 | 169 | 2 | 55 |
AP2 2 gykGVrwdkkrgrWvAeIrdpseng.krkrfslgkfgtaeeAakaaiaarkkleg 55
+y+GVr+++ +g+++AeIrd k +r++lg+f+taeeAa a+++a+++++g
WALNUT_00015337-RA 119 HYRGVRRRP-WGKYAAEIRDS---SrKGARVWLGTFDTAEEAALAYDKAALRIRG 169
7********.**********3...2234*************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:3.30.730.10 | 6.5E-32 | 118 | 178 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 5.8E-38 | 119 | 183 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 1.52E-30 | 119 | 177 | No hit | No description |
| SuperFamily | SSF54171 | 9.15E-24 | 119 | 179 | IPR016177 | DNA-binding domain |
| PROSITE profile | PS51032 | 24.092 | 119 | 177 | IPR001471 | AP2/ERF domain |
| Pfam | PF00847 | 4.0E-15 | 119 | 169 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 2.6E-11 | 120 | 131 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 2.6E-11 | 143 | 159 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 279 aa Download sequence Send to blast |
MSKNCEDPPA PGAILENVWA SFISGDGPGT TYTPELSRPW EELPSLDRRD GSKEILRRLP 60 SLGRWISMGA EAWEALLDGI VVPASNMEQS CNDNLECNGA TGLCLQKPNA KAEKAATRHY 120 RGVRRRPWGK YAAEIRDSSR KGARVWLGTF DTAEEAALAY DKAALRIRGP KAYLNFPLET 180 VAEALGLDYS GNDVSCSSMS NKGLGEDPAC TFLGCSAEKV CNPRKRSASR EFEQDIDMII 240 EQPALKKTRS MENILGNEFD VVEFQDLGTD YLDSLLSSF |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2gcc_A | 8e-28 | 115 | 180 | 1 | 66 | ATERF1 |
| 3gcc_A | 8e-28 | 115 | 180 | 1 | 66 | ATERF1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00441 | DAP | Transfer from AT4G18450 | Download |
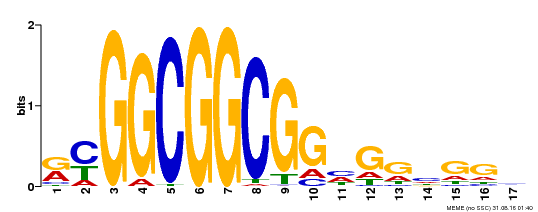
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018838470.1 | 0.0 | PREDICTED: ethylene-responsive transcription factor ERF091-like | ||||
| TrEMBL | A0A2I4G3M8 | 0.0 | A0A2I4G3M8_JUGRE; ethylene-responsive transcription factor ERF091-like | ||||
| STRING | EOY28113 | 1e-111 | (Theobroma cacao) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF9547 | 31 | 42 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G18450.1 | 2e-42 | ERF family protein | ||||



