 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | WALNUT_00014608-RA | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fagales; Juglandaceae; Juglans
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 556aa MW: 60509.4 Da PI: 4.778 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 54 | 3.8e-17 | 39 | 86 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT Ed +lvd vk++G g+W+++ ++ g+ R++k+c++rw ++l
WALNUT_00014608-RA 39 KGPWTSTEDAILVDFVKKHGEGNWNAVQKHSGLFRCGKSCRLRWANHL 86
79******************************************9986 PP
| |||||||
| 2 | Myb_DNA-binding | 51.7 | 1.9e-16 | 92 | 135 | 1 | 46 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqk 46
+g++T+eE+ l++++++++G++ W++ a++++ gRt++++k++w++
WALNUT_00014608-RA 92 KGAFTAEEERLIIELHAKMGNK-WARMAAHLP-GRTDNEIKNYWNT 135
799*******************.*********.***********96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 15.641 | 34 | 86 | IPR017930 | Myb domain |
| SMART | SM00717 | 1.2E-14 | 38 | 88 | IPR001005 | SANT/Myb domain |
| SuperFamily | SSF46689 | 6.74E-30 | 38 | 133 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 7.8E-15 | 39 | 86 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 4.5E-23 | 40 | 93 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 2.66E-11 | 41 | 86 | No hit | No description |
| PROSITE profile | PS51294 | 26.42 | 87 | 141 | IPR017930 | Myb domain |
| SMART | SM00717 | 3.8E-17 | 91 | 139 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.9E-15 | 92 | 135 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 8.5E-26 | 94 | 140 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 1.39E-12 | 94 | 135 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009789 | Biological Process | positive regulation of abscisic acid-activated signaling pathway | ||||
| GO:0043068 | Biological Process | positive regulation of programmed cell death | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0045926 | Biological Process | negative regulation of growth | ||||
| GO:0048235 | Biological Process | pollen sperm cell differentiation | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 556 aa Download sequence Send to blast |
MPCKTTESEE GSLPKDQCES PLINENNEGC AGGGVVLKKG PWTSTEDAIL VDFVKKHGEG 60 NWNAVQKHSG LFRCGKSCRL RWANHLRPNL KKGAFTAEEE RLIIELHAKM GNKWARMAAH 120 LPGRTDNEIK NYWNTRIKRR QRAGLPLYPP EVCFQALQES QHGQNTVGVN GGDRGHHDIL 180 LTNSYEIPDV IFDSLKANQG VLPYVPELPD IPTSSMLLKG LGSPQYCTLQ PIMPIQKRLR 240 ESNELFPGPS ASVGNGFSSF EQFQDDTCDK VALSFGFSLP LDPDPAIKNS ESFGVIQGCH 300 SLSNGNSSAS KPTSRAVKLE LPSLQYAETD FSNWVASSPR PLLESVDAFI KSPPPAGTFE 360 IDCSSPRNSG LLDALLQESK NLSSSKKHSS DKSSNSSSAT PGDIADSSTL NICETEWADY 420 GDPISPFGHS TTSLFSEISP ISASGSSMDG QPSAETFTGI IVKSEPVDIA WTSDREEEST 480 ARLSFTQPDA LLGVDWLEQS SGCIKDQSTF ATFLDDDFDI DYKHMAAGTS TSNQGWGLNS 540 FAWNNMPAVC QMSDLP |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 2e-32 | 37 | 140 | 5 | 107 | B-MYB |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00490 | DAP | Transfer from AT5G06100 | Download |
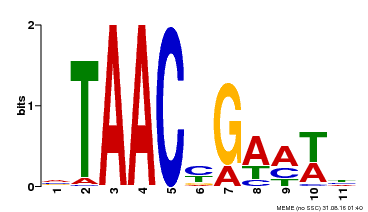
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018849363.1 | 0.0 | PREDICTED: transcription factor GAMYB | ||||
| Refseq | XP_018849365.1 | 0.0 | PREDICTED: transcription factor GAMYB | ||||
| Refseq | XP_018849366.1 | 0.0 | PREDICTED: transcription factor GAMYB | ||||
| Refseq | XP_018849367.1 | 0.0 | PREDICTED: transcription factor GAMYB | ||||
| TrEMBL | A0A2I4GZM8 | 0.0 | A0A2I4GZM8_JUGRE; transcription factor GAMYB | ||||
| STRING | XP_008231582.1 | 0.0 | (Prunus mume) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF2047 | 34 | 85 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G06100.3 | 1e-132 | myb domain protein 33 | ||||



