 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | WALNUT_00010733-RA | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fagales; Juglandaceae; Juglans
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 457aa MW: 52069.3 Da PI: 6.0056 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 57.3 | 3.5e-18 | 211 | 257 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g WT++Ed ll+++v ++G++ W+ Iar+++ gR +kqc++rw+++l
WALNUT_00010733-RA 211 KGQWTPQEDRLLLQLVDRFGTKKWSHIARMLN-GRMGKQCRERWHNHL 257
799*****************************.*************97 PP
| |||||||
| 2 | Myb_DNA-binding | 58 | 2.2e-18 | 263 | 305 | 1 | 45 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwq 45
+++W++eEd++l++a+k+ G++ W++Iar+++ gRt++ +k++w+
WALNUT_00010733-RA 263 KDAWSEEEDKILIEAHKEIGNK-WAAIARRLP-GRTENTIKNHWN 305
679*******************.*********.***********8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 30.647 | 206 | 261 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 6.24E-32 | 208 | 304 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 2.6E-17 | 210 | 259 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.3E-24 | 212 | 261 | IPR009057 | Homeodomain-like |
| Pfam | PF13921 | 2.0E-18 | 214 | 271 | No hit | No description |
| CDD | cd00167 | 3.83E-15 | 214 | 257 | No hit | No description |
| PROSITE profile | PS51294 | 21.298 | 262 | 312 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 3.7E-22 | 262 | 312 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 2.5E-16 | 262 | 310 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 9.15E-13 | 265 | 308 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010439 | Biological Process | regulation of glucosinolate biosynthetic process | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:1904095 | Biological Process | negative regulation of endosperm development | ||||
| GO:2000692 | Biological Process | negative regulation of seed maturation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 457 aa Download sequence Send to blast |
MEFDANFRED FPFLSNLFSD NNTPPKPELT DGFSLEPSAS CKGNLFDDLH GHFDHFPLSG 60 SNLNPYDHDG DHFNVEGSTS NPLSGIRECP CTDPCEAYAN AFLNDFTASL FASDSTNGNM 120 QGLESSYLSF WDNPDQKNLV QPTTGGQSHI YLPLNIQEHD GPTNGRFADD ISCITAENEY 180 NRKLAVQKKI RKRAVHMRKA SEVQKKPSII KGQWTPQEDR LLLQLVDRFG TKKWSHIARM 240 LNGRMGKQCR ERWHNHLRPD IRKDAWSEEE DKILIEAHKE IGNKWAAIAR RLPGRTENTI 300 KNHWNATKRR QLSKKNRKDS DSNGSLLQSY IKSVTSASAP RIDKNINMLS ASGKQMVSMM 360 TKPRIDQLES SDHLNSADWA IPVYNDHNYY EAMGLLFDKN MLPETLGFMS MLEEMPCSSL 420 VEESNIEYET PLETDNLVQG EAKKEMDLME LIYQGKL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 1e-44 | 207 | 312 | 3 | 108 | B-MYB |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 181 | 192 | RKLAVQKKIRKR |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00385 | DAP | Transfer from AT3G27785 | Download |
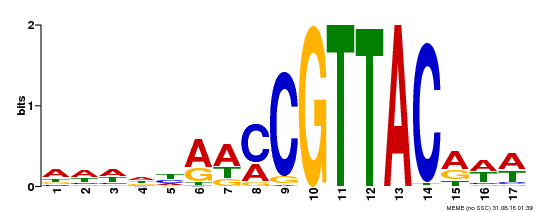
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018822991.1 | 0.0 | PREDICTED: transcription factor MYB98-like | ||||
| TrEMBL | A0A2I4EUB7 | 0.0 | A0A2I4EUB7_JUGRE; transcription factor MYB98-like | ||||
| STRING | EOY02441 | 1e-138 | (Theobroma cacao) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1570 | 31 | 96 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G27785.1 | 4e-73 | myb domain protein 118 | ||||



