 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Jcr4S05562.10 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Euphorbiaceae; Crotonoideae; Jatropheae; Jatropha
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 1042aa MW: 115999 Da PI: 6.0465 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 54.7 | 1.7e-17 | 106 | 164 | 3 | 57 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHC....TS-HHHHHHHHHHHHHHHHC CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkkl....gLterqVkvWFqNrRakekk 57
k ++t+eq+e+Le+l++++++ps +r++ +++ +++ +q+kvWFqNrR +ek+
Jcr4S05562.10 106 KYVRYTPEQVEALERLYHECPKPSSIRRQQFIRECpilsNIEPKQIKVWFQNRRCREKQ 164
5679*****************************************************97 PP
| |||||||
| 2 | START | 179.9 | 1.5e-56 | 250 | 458 | 2 | 205 |
HHHHHHHHHHHHHHHC-TT-EEEEEXCCTTEEEEEEESSS.SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-SEEEEEEEECTT..EEEEEEE CS
START 2 laeeaaqelvkkalaeepgWvkssesengdevlqkfeeskvdsgealrasgvvdmvlallveellddkeqWdetlakaetlevissg..galqlmv 95
+aee+++e+++ka+ ++ Wv+++ +++g++++ +++ s++++g a+ra+g+v +++ v+e+l+d++ W ++++++++l+v+ ++ g+++l +
Jcr4S05562.10 250 IAEETLTEFLSKATGTAVEWVQMPGMKPGPDSIGIIAISHGCTGVAARACGLVGLEPT-RVAEILKDRPSWFRDCRAVDVLNVLPTAngGTIELLY 344
789*******************************************************.8888888888****************9999******* PP
EXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--....-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE--SSXXHHHHHHHHHH CS
START 96 aelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe...sssvvRaellpSgiliepksnghskvtwvehvdlkgrlphwllrslvks 187
++l+a+++l+p Rdf+ +Ry+ l++g++vi+++S+ ++q+ p+ +++vRae+lpSg+li+p+++g+s +++v+h+dl+ ++++++lr+l++s
Jcr4S05562.10 345 MQLYAPTTLAPaRDFWLLRYTSVLEDGSLVICERSLKNTQNGPSmppVQHFVRAEMLPSGYLIRPCEGGGSIIHIVDHMDLEPWSVPEVLRPLYES 440
******************************************9988899*********************************************** PP
HHHHHHHHHHHHTXXXXX CS
START 188 glaegaktwvatlqrqce 205
+++ ++kt++a l+++++
Jcr4S05562.10 441 STVLAQKTTMAGLRQLRQ 458
*************99876 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50071 | 15.111 | 101 | 165 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 5.7E-14 | 103 | 169 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 1.24E-15 | 105 | 169 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 3.23E-15 | 106 | 166 | No hit | No description |
| Pfam | PF00046 | 4.2E-15 | 107 | 164 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 1.2E-17 | 108 | 164 | IPR009057 | Homeodomain-like |
| CDD | cd14686 | 1.56E-6 | 158 | 197 | No hit | No description |
| PROSITE profile | PS50848 | 25.388 | 240 | 455 | IPR002913 | START domain |
| CDD | cd08875 | 5.86E-79 | 244 | 460 | No hit | No description |
| SMART | SM00234 | 3.2E-42 | 249 | 459 | IPR002913 | START domain |
| Gene3D | G3DSA:3.30.530.20 | 1.2E-23 | 249 | 456 | IPR023393 | START-like domain |
| SuperFamily | SSF55961 | 1.02E-37 | 249 | 460 | No hit | No description |
| Pfam | PF01852 | 5.6E-54 | 250 | 458 | IPR002913 | START domain |
| Pfam | PF08670 | 2.7E-9 | 793 | 843 | IPR013978 | MEKHLA |
| Pfam | PF08670 | 3.1E-32 | 888 | 988 | IPR013978 | MEKHLA |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0010014 | Biological Process | meristem initiation | ||||
| GO:0010075 | Biological Process | regulation of meristem growth | ||||
| GO:0010087 | Biological Process | phloem or xylem histogenesis | ||||
| GO:0048263 | Biological Process | determination of dorsal identity | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1042 aa Download sequence Send to blast |
MIVSEAWIGS TVPFWLVKEE ISYLITNQWF MLSCVLGLSN NRQNEDNDVS HLDIAAPVSF 60 LSGILKLLIK PLESYLALRK ERVKLNFKMA MSCKDGKQPA NLDNGKYVRY TPEQVEALER 120 LYHECPKPSS IRRQQFIREC PILSNIEPKQ IKVWFQNRRC REKQRKEASR LQAVNRKLTA 180 MNKLLMEEND RLQKQVSQLV YENGHFRQHT QNTTLATKDT SCESVVTSGQ HHLTPQHPPR 240 DASPAGLLSI AEETLTEFLS KATGTAVEWV QMPGMKPGPD SIGIIAISHG CTGVAARACG 300 LVGLEPTRVA EILKDRPSWF RDCRAVDVLN VLPTANGGTI ELLYMQLYAP TTLAPARDFW 360 LLRYTSVLED GSLVICERSL KNTQNGPSMP PVQHFVRAEM LPSGYLIRPC EGGGSIIHIV 420 DHMDLEPWSV PEVLRPLYES STVLAQKTTM AGLRQLRQIA QEVSQSNVTN WGRRPAALRA 480 LSQRGFNEAL NGFTDEGWSM MGNDGMDDVT ILVNSSPEKL MGLNLSFTNG FPSVSNAVLC 540 AKASMLLQNV PPAILLRFLR EHRSEWADNN IDAYSAAAIK IGPCSLPGAR VGSFGGQVIL 600 PLAHTIEHEE FLEVIKLEGV GHSPEDPIMP RDMFLLQYAQ SFNDNWLSLL CSGMDENAIG 660 TCAELIFAPI DASFADDAPL LPSGFRIIPL DSAKEASSPN RTLDLASALE IGPAGNKSST 720 EYSANSGCVR SVMTIAFEFA FESHMQEHVA SMARQYVRSI ISSVQRVALA LSPSHLGSHA 780 GLRPPLGTPE AQTLARWICL SYRCYLGVEL LKSSSEGSES VLKTLWHHSD AIMCCSPEGP 840 YYFFSLYFFF LQWILEMEIL KICSDYASYN GVVYVLEIKE IGVFSNCYVF RCEFQALPVF 900 TFANQAGLDM LETTLVALQD ITLEKIFDDH GRKTLCSEFP QIMQQGFACL QGGICLSSMG 960 RPVSYERAVA WKVLNEEENA HCICFILAEL NEGAALLLII QMRLNDIDLK SLYPLKGAPR 1020 VALQFLFAFI FRSGQVLEYD AK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of meristem development to promote lateral organ formation. May regulates procambial and vascular tissue formation or maintenance, and vascular development in inflorescence stems. {ECO:0000269|PubMed:15598805, ECO:0000269|PubMed:15705957, ECO:0000269|PubMed:16617092}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
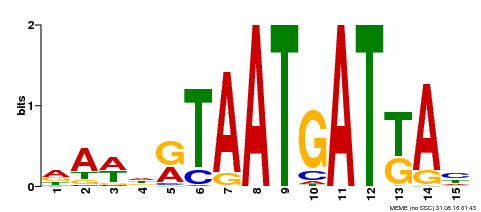
| Motif ID | Method | Source | Motif file |
| MP00200 | DAP | Transfer from AT1G52150 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin. Repressed by miR165 and miR166. {ECO:0000269|PubMed:15773855, ECO:0000269|PubMed:16033795, ECO:0000269|PubMed:16617092, ECO:0000269|PubMed:17237362}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KF534716 | 0.0 | KF534716.1 Populus alba x Populus glandulosa HD-ZIPIII mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_012069098.1 | 0.0 | homeobox-leucine zipper protein ATHB-15 | ||||
| Swissprot | Q9ZU11 | 0.0 | ATB15_ARATH; Homeobox-leucine zipper protein ATHB-15 | ||||
| TrEMBL | A0A067KXE1 | 0.0 | A0A067KXE1_JATCU; Uncharacterized protein | ||||
| STRING | XP_002515977.1 | 0.0 | (Ricinus communis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1946 | 33 | 87 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G52150.1 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



