 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Jcr4S02818.20 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Euphorbiaceae; Crotonoideae; Jatropheae; Jatropha
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 860aa MW: 98908.7 Da PI: 7.5151 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 59.7 | 9.2e-19 | 111 | 198 | 16 | 91 |
FAR1 16 kskskkskrngeitkrtfvCskegkreeekkk............tekerrtraetrtgCkaklkvkkekdgkwevtkleleHnHelap 91
+++s++sk+++e+++++f Cs++g+++e +k+ e+ +r+ ++t+Cka+++vk++ dgkw+++++++eHnH+l p
Jcr4S02818.20 111 IQNSRRSKTSREFIDAKFACSRYGTKREYDKSfnrprarqnkqdPENGTGRRSCSKTDCKASMHVKRRPDGKWVIHSFVKEHNHDLLP 198
6789999**********************999999998776555444555999********************************975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 7.4E-17 | 111 | 198 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 4.2E-27 | 296 | 387 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 9.532 | 576 | 612 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 3.2E-6 | 587 | 614 | IPR006564 | Zinc finger, PMZ-type |
| Pfam | PF04434 | 9.7E-5 | 588 | 611 | IPR007527 | Zinc finger, SWIM-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009585 | Biological Process | red, far-red light phototransduction | ||||
| GO:0010218 | Biological Process | response to far red light | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 860 aa Download sequence Send to blast |
MEAIVAIYSV NHFFINFSSL LHGTRLVLAK RKNLSFMDID LRLPSGDHDK DNEEPSGIDN 60 MLTEEKLHNG DVATGSIVDV AEEVHAIEGG HMSSPTTEFK EEANLEPLSA IQNSRRSKTS 120 REFIDAKFAC SRYGTKREYD KSFNRPRARQ NKQDPENGTG RRSCSKTDCK ASMHVKRRPD 180 GKWVIHSFVK EHNHDLLPAQ AVSEQTRKMY AAMARQFAEY KHVVGLKNDP KNPFDKGRNL 240 ALEAADAKIL LDFFTQMQNL NSNFFYAIEL GEDQRLKNLF WVDAKSRHDY VNFSDVVSFD 300 TIYVRNKYKM PLALFVGVNQ HYQFMLLGCA LLSDENATTY SWLMQTWLRA MGGQAPKVII 360 TDQDKALKSV ISEVLPNAHH YFFLWNILGK VSENLSQVIK QHENFIPKFE KCIFRSWTND 420 EFVKRWLKIL DRFELRENEL MQSLYEDREL WVPIYMRDAI LAGMSMTQRS ESINSYFDKY 480 VHKKTTVQEF VKQYETILQD RYEEEAKADS DTWNKQPTLK SPSPLEKSVS GVYTHAVFKK 540 FQVEVLGVVA CHPKMESQDE TSISFRVQDL EKHQDYTVVW NQIRSEVACI CRLYEYKGYL 600 CRHALVVLQM CQQSAIPPQY ILKRWTKDVK NRHFFGEESE QVQSRFQRYN ELCQRALKLS 660 EEGSLSQESY NIAFRALEEA FGNCISANNS SKTLAEAGTA ATHGLLCIEE DNQNRSMNKT 720 NKKKNPTKKR KVNSEQEVTT LGAEDSLQQM DKLNSRSVTL DGYYGAQQSV PGMVQLNLMA 780 PTRDNYYGNQ QTIQGLGQLN SIAPSHDGYY NAQQSMHGLG QMDFFRAQAG FTYGIRVSWL 840 DVDDPNVRTA PLHDNASRHA |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. When associated with PHYA, protects it from being recognized and degraded by the COP1/SPA complex. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:17012604, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:18715961, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00078 | ChIP-seq | Transfer from AT3G22170 | Download |
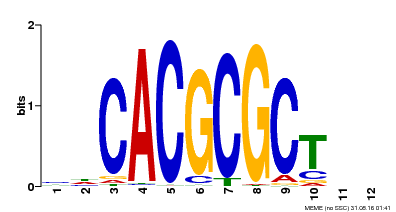
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. Up-regulated by white light. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_012092528.1 | 0.0 | protein FAR-RED ELONGATED HYPOCOTYL 3 isoform X2 | ||||
| Swissprot | Q9LIE5 | 0.0 | FHY3_ARATH; Protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| TrEMBL | A0A067JCL9 | 0.0 | A0A067JCL9_JATCU; Uncharacterized protein | ||||
| STRING | POPTR_0006s02150.1 | 0.0 | (Populus trichocarpa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF40 | 30 | 530 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G22170.2 | 0.0 | far-red elongated hypocotyls 3 | ||||



