 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Jcr4S02286.50 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Euphorbiaceae; Crotonoideae; Jatropheae; Jatropha
|
||||||||
| Family | MIKC_MADS | ||||||||
| Protein Properties | Length: 256aa MW: 29128.2 Da PI: 7.8881 | ||||||||
| Description | MIKC_MADS family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SRF-TF | 89.6 | 1.6e-28 | 9 | 59 | 1 | 51 |
S---SHHHHHHHHHHHHHHHHHHHHHHHHHHT-EEEEEEE-TTSEEEEEE- CS
SRF-TF 1 krienksnrqvtfskRrngilKKAeELSvLCdaevaviifsstgklyeyss 51
k+ien rqvtfskRr g++KKA+ELSvLCdaevavi+fsstgklye+ss
Jcr4S02286.50 9 KKIENLNSRQVTFSKRRSGLIKKAKELSVLCDAEVAVIVFSSTGKLYEFSS 59
68***********************************************96 PP
| |||||||
| 2 | K-box | 67 | 6.4e-23 | 90 | 176 | 14 | 100 |
K-box 14 aeslqqelakLkkeienLqreqRhllGedLesLslkeLqqLeqqLekslkkiRskKnellleqieelqkkekelqeenkaLrkklee 100
++s ++e++ Lk+e+++L+ + +++G++L+ Ls+keLqqLe+qL ++ ++++kK++++leq+++++ +e+++ +en++Lrk++ee
Jcr4S02286.50 90 EQSQSEEVTALKDEVSKLRLTCLQMMGQQLDGLSFKELQQLEHQLTEGRISVKEKKEKVILEQLKKSRLQEQKAIQENETLRKQVEE 176
68899*******************************************************************************987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00432 | 8.7E-42 | 1 | 60 | IPR002100 | Transcription factor, MADS-box |
| PROSITE profile | PS50066 | 32.842 | 1 | 61 | IPR002100 | Transcription factor, MADS-box |
| CDD | cd00265 | 9.58E-38 | 2 | 61 | No hit | No description |
| SuperFamily | SSF55455 | 6.28E-30 | 2 | 65 | IPR002100 | Transcription factor, MADS-box |
| PROSITE pattern | PS00350 | 0 | 3 | 57 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 2.1E-29 | 3 | 23 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF00319 | 3.7E-26 | 10 | 57 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 2.1E-29 | 23 | 38 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 2.1E-29 | 38 | 59 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF01486 | 4.9E-17 | 90 | 174 | IPR002487 | Transcription factor, K-box |
| PROSITE profile | PS51297 | 13.18 | 90 | 180 | IPR002487 | Transcription factor, K-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010047 | Biological Process | fruit dehiscence | ||||
| GO:0010262 | Biological Process | somatic embryogenesis | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0048577 | Biological Process | negative regulation of short-day photoperiodism, flowering | ||||
| GO:0060862 | Biological Process | negative regulation of floral organ abscission | ||||
| GO:0060867 | Biological Process | fruit abscission | ||||
| GO:0071365 | Biological Process | cellular response to auxin stimulus | ||||
| GO:2000692 | Biological Process | negative regulation of seed maturation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005737 | Cellular Component | cytoplasm | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 256 aa Download sequence Send to blast |
MGRGKIEIKK IENLNSRQVT FSKRRSGLIK KAKELSVLCD AEVAVIVFSS TGKLYEFSSS 60 SLTWNKLCQD IAKARILMCP EHASDYPGTE QSQSEEVTAL KDEVSKLRLT CLQMMGQQLD 120 GLSFKELQQL EHQLTEGRIS VKEKKEKVIL EQLKKSRLQE QKAIQENETL RKQVEELKRS 180 SAMAKSQFQE FNPLERRFPL ASFKSDCFPQ GEEEDYGDLS ETSLHLGLSY DVHHKRKAAK 240 IEPIWNDSAG SQVASE |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5f28_A | 2e-18 | 1 | 62 | 1 | 62 | MEF2C |
| 5f28_B | 2e-18 | 1 | 62 | 1 | 62 | MEF2C |
| 5f28_C | 2e-18 | 1 | 62 | 1 | 62 | MEF2C |
| 5f28_D | 2e-18 | 1 | 62 | 1 | 62 | MEF2C |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00508 | DAP | Transfer from AT5G13790 | Download |
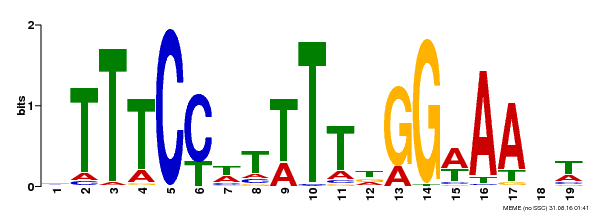
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP011969 | 2e-96 | AP011969.1 Jatropha curcas DNA, clone: JHL06B08, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_012086200.1 | 1e-173 | MADS-box transcription factor 23 isoform X2 | ||||
| TrEMBL | E6NU98 | 1e-172 | E6NU98_JATCU; JHL06B08.10 protein | ||||
| STRING | POPTR_0006s04730.1 | 2e-98 | (Populus trichocarpa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF3582 | 32 | 60 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G13790.1 | 2e-60 | AGAMOUS-like 15 | ||||



