 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Jcr4S01139.30 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Euphorbiaceae; Crotonoideae; Jatropheae; Jatropha
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 909aa MW: 99922.6 Da PI: 6.5478 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 66.2 | 4.5e-21 | 116 | 171 | 1 | 56 |
TT--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 1 rrkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ +++t++q++eLe+lF+++++p++++r eL+k+l+L++rqVk+WFqNrR+++k
Jcr4S01139.30 116 KKRYHRHTPQQIQELEALFKECPHPDEKQRLELSKRLSLETRQVKFWFQNRRTQMK 171
688999***********************************************999 PP
| |||||||
| 2 | START | 145 | 7.2e-46 | 325 | 515 | 1 | 172 |
HHHHHHHHHHHHHHHHC-TT-EEEE....EXCCTTEEEEEEESSS......SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-S....EEEEEE CS
START 1 elaeeaaqelvkkalaeepgWvkss....esengdevlqkfeeskv.....dsgealrasgvvdmvlallveellddkeqWdetla....kaetle 83
ela++a++elvk+a+ +ep+W +s e++n +e++++f++ + + +ea+r++g v+ ++ lve+l+d++ +W e+++ + +t++
Jcr4S01139.30 325 ELALAAMDELVKMAQTDEPLWIRSLeggrEILNHEEYMRTFTPCIGmkpsgFFSEASRETGTVIINSLALVETLMDSN-RWAEMFPcmiaRTTTTD 419
5899**************************************99989*******************************.***************** PP
EECTT......EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--.-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE- CS
START 84 vissg......galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppesssvvRaellpSgiliepksnghskvtwvehvdl 172
vissg g lqlm aelq+lsplvp R++ f+R+++q+ +g+w++vdvS+d ++ + + ++ +++lpSg+++++++ng+sk +v dl
Jcr4S01139.30 420 VISSGmggtrnGSLQLMHAELQVLSPLVPvREVNFLRFCKQHAEGVWAVVDVSIDTIRETSGAPTFINCRRLPSGCVVQDMPNGYSKPEYVLTRDL 515
***********************************************************999****************************998776 PP
| |||||||
| 3 | START | 44.8 | 3.4e-15 | 561 | 603 | 164 | 206 |
EEEEE-EE--SSXXHHHHHHHHHHHHHHHHHHHHHHTXXXXXX CS
START 164 vtwvehvdlkgrlphwllrslvksglaegaktwvatlqrqcek 206
vtwveh++++++++h+l+r+l++sg+ +ga++wvatlqrqce+
Jcr4S01139.30 561 VTWVEHAEYEESQIHQLYRPLISSGMGFGAQRWVATLQRQCEC 603
8****************************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.09E-20 | 99 | 173 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 1.1E-21 | 102 | 167 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.605 | 113 | 173 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 2.4E-19 | 114 | 177 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 4.53E-19 | 115 | 173 | No hit | No description |
| Pfam | PF00046 | 1.3E-18 | 116 | 171 | IPR001356 | Homeobox domain |
| PROSITE pattern | PS00027 | 0 | 148 | 171 | IPR017970 | Homeobox, conserved site |
| PROSITE profile | PS50848 | 38.521 | 316 | 606 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 5.16E-29 | 318 | 503 | No hit | No description |
| CDD | cd08875 | 4.25E-117 | 320 | 602 | No hit | No description |
| Pfam | PF01852 | 1.8E-37 | 325 | 515 | IPR002913 | START domain |
| SMART | SM00234 | 1.9E-38 | 325 | 603 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 5.16E-29 | 558 | 603 | No hit | No description |
| Pfam | PF01852 | 2.0E-11 | 561 | 603 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 1.08E-18 | 631 | 726 | No hit | No description |
| SuperFamily | SSF55961 | 1.08E-18 | 768 | 902 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009827 | Biological Process | plant-type cell wall modification | ||||
| GO:0042335 | Biological Process | cuticle development | ||||
| GO:0043481 | Biological Process | anthocyanin accumulation in tissues in response to UV light | ||||
| GO:0048765 | Biological Process | root hair cell differentiation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 909 aa Download sequence Send to blast |
MSFGGFLENG SPGGGGARIV ADIPYSSSNM PTGAIAQPRL VSPSLTKSMF SSPGLSLALQ 60 QPNIDSPGDM GRMAENFEPS GGRRSREEEH ESRSGSDNMD GASGDDQDAA DNPPRKKRYH 120 RHTPQQIQEL EALFKECPHP DEKQRLELSK RLSLETRQVK FWFQNRRTQM KTQLERHENS 180 LLRQENDKLR AENMSIRDAM RNPICSNCGG PAIIGDISLE EQHLRIENAR LKDELDRVCA 240 LAGKFLGRPI SSLAGSIGPP MPNSSLELGV GSNGFGGLST VATTLPLGPD FGGGISSLPV 300 MNQPRSTTTG VTGLDRSLER SMFLELALAA MDELVKMAQT DEPLWIRSLE GGREILNHEE 360 YMRTFTPCIG MKPSGFFSEA SRETGTVIIN SLALVETLMD SNRWAEMFPC MIARTTTTDV 420 ISSGMGGTRN GSLQLMHAEL QVLSPLVPVR EVNFLRFCKQ HAEGVWAVVD VSIDTIRETS 480 GAPTFINCRR LPSGCVVQDM PNGYSKPEYV LTRDLIVKLH DCAILFPYEY PNPFLNNSTH 540 LDVAFYVPRA IYEICLHFPL VTWVEHAEYE ESQIHQLYRP LISSGMGFGA QRWVATLQRQ 600 CECLAILMSS TVPSRDHTAI TASGRRSMLK LAQRMTDNFC AGVCASTVHK WNKLNAGNVD 660 EDVRVMTRKS VDDPGEPPGI VLSAATSVWL PVSPQRLFDF LRDERLRSEW DILSKRRTHA 720 GDGPHSQRPG SRQLRLAVTC QRELERTSWE FGIIQMDRHR QTVVALREIY HIILSVQAMN 780 ANQSSMLILQ ETCIDAAGSL VVYAPVDIPA MHVVMNGGDS AYVALLPSGF SIVPDGPGSR 840 GSPSTNANGP SSNNGGGQQR VSGSLLTVAF QILVNSLPTA KLTVESVETV NNLISCTVQK 900 IKAALQCES |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 714 | 718 | KRRTH |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of the tissue-specific accumulation of anthocyanins and in cellular organization of the primary root. {ECO:0000269|PubMed:10402424}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00421 | DAP | Transfer from AT4G00730 | Download |
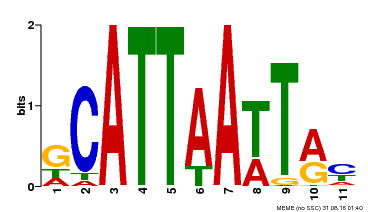
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_012083470.1 | 0.0 | homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| Swissprot | Q0WV12 | 0.0 | ANL2_ARATH; Homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| TrEMBL | A0A067JXS1 | 0.0 | A0A067JXS1_JATCU; Uncharacterized protein | ||||
| STRING | cassava4.1_001764m | 0.0 | (Manihot esculenta) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1192 | 34 | 106 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G00730.1 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



