 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Jcr4S00077.70 | ||||||||
| Common Name | JCGZ_24594, LOC105631288 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Euphorbiaceae; Crotonoideae; Jatropheae; Jatropha
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 578aa MW: 62680.6 Da PI: 5.0112 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 56.4 | 6.7e-18 | 40 | 87 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT Ed +l+++vk++G g+W+++ ++ g+ R++k+c++rw ++l
Jcr4S00077.70 40 KGPWTSAEDAILIEYVKKHGEGNWNAVQKHSGLSRCGKSCRLRWANHL 87
79******************************************9986 PP
| |||||||
| 2 | Myb_DNA-binding | 40.3 | 7.5e-13 | 93 | 157 | 1 | 46 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHT....................TTS-HHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmg....................kgRtlkqcksrwqk 46
+g++T+eE++l++++++++G++ W++ a++++ +gRt++++k++w++
Jcr4S00077.70 93 KGAFTQEEEQLIIELHAKMGNK-WARMAAHIPefksrlvpmetfisfmytscLGRTDNEIKNYWNT 157
799*******************.*****************************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 23.548 | 35 | 91 | IPR017930 | Myb domain |
| SMART | SM00717 | 3.6E-16 | 39 | 89 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 4.4E-16 | 40 | 87 | IPR001005 | SANT/Myb domain |
| SuperFamily | SSF46689 | 4.3E-25 | 41 | 117 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 8.0E-25 | 41 | 94 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 3.66E-12 | 42 | 87 | No hit | No description |
| PROSITE profile | PS51294 | 15.61 | 92 | 163 | IPR017930 | Myb domain |
| SMART | SM00717 | 9.3E-13 | 92 | 161 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 3.8E-12 | 93 | 157 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 3.72E-8 | 95 | 157 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 4.5E-23 | 95 | 162 | IPR009057 | Homeodomain-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009789 | Biological Process | positive regulation of abscisic acid-activated signaling pathway | ||||
| GO:0043068 | Biological Process | positive regulation of programmed cell death | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0045926 | Biological Process | negative regulation of growth | ||||
| GO:0048235 | Biological Process | pollen sperm cell differentiation | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 578 aa Download sequence Send to blast |
MSHTTSESDD GVLSKDRADS PLAEEGSCGG SANGGIVLKK GPWTSAEDAI LIEYVKKHGE 60 GNWNAVQKHS GLSRCGKSCR LRWANHLRPN LKKGAFTQEE EQLIIELHAK MGNKWARMAA 120 HIPEFKSRLV PMETFISFMY TSCLGRTDNE IKNYWNTRIK RRQRAGLPLY PPEVSLQALQ 180 ESQRGLNVGG ISTGDKGHQD LMRTNSYEIP DVIFDSLKAN QGISPYVSEL PDISASSILV 240 KGLGSSQYGS FMSPSVHRQK RFRESTTLIP GYSGGVKTEF PLFDQFQDNS CDKVAQSFGL 300 SFPFDPDPTN KNPQSFCDNQ GGHTLANGDF SASKPNSGPL KMELPSLQYP ENDLGSWGTS 360 PPHLLETVDT FIQSPPTGAV ESSPRNSGLL DALLHEAKTL SSAKNHSSDK SSNSSTVTPG 420 ELAESSALNI CGAEWEDYGD PLSPLGHTAT SLFSDCTPIS ASGSSLDEPP PTETFTGCNV 480 KSELVDQAWS PERERETTNR LDITRPDALL ASDWLGQGSG YVKEQVIMAD NIATLLGDDL 540 SSDYKQISAG ASTSNQGWGL GSCAWNNMPA VCQMSELP |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 5e-28 | 38 | 162 | 5 | 107 | B-MYB |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00490 | DAP | Transfer from AT5G06100 | Download |
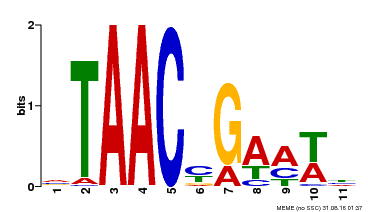
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KM034648 | 0.0 | KM034648.1 Jatropha curcas clone JcMYB028 MYB family protein gene, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_012068750.1 | 0.0 | transcription factor GAMYB | ||||
| Refseq | XP_012068751.1 | 0.0 | transcription factor GAMYB | ||||
| Refseq | XP_020534006.1 | 0.0 | transcription factor GAMYB | ||||
| TrEMBL | A0A067L7W6 | 0.0 | A0A067L7W6_JATCU; MYB family protein | ||||
| STRING | XP_002525574.1 | 0.0 | (Ricinus communis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF2047 | 34 | 85 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G06100.3 | 1e-143 | myb domain protein 33 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Entrez Gene | 105631288 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



