 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Jcr4S00070.110 | ||||||||
| Common Name | JCGZ_12574, LOC105639879 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Euphorbiaceae; Crotonoideae; Jatropheae; Jatropha
|
||||||||
| Family | AP2 | ||||||||
| Protein Properties | Length: 661aa MW: 73334.1 Da PI: 6.959 | ||||||||
| Description | AP2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 39.1 | 1.9e-12 | 322 | 378 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrd.pse.ng.krkrfslgkfgtaeeAakaaiaarkkleg 55
s+y+GV++++++gr++A+++d ++ g rk + g ++ +e+Aa+a++ a++k++g
Jcr4S00070.110 322 SQYRGVTRHRWTGRYEAHLWDnSCKkEGqSRKGRQ-GGYDMEEKAARAYDLAALKYWG 378
78*******************87776664446655.77******************98 PP
| |||||||
| 2 | AP2 | 45.2 | 2.3e-14 | 423 | 472 | 3 | 55 |
AP2 3 ykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
y+GV+++++ grW A+I + +k +lg+f t eeAa+a++ a+ k++g
Jcr4S00070.110 423 YRGVTRHHQHGRWQARIGRVAG---NKDLYLGTFSTQEEAAEAYDVAAIKFRG 472
9***************988532...5************************997 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00018 | 1.61E-18 | 322 | 386 | No hit | No description |
| SuperFamily | SSF54171 | 5.23E-14 | 322 | 387 | IPR016177 | DNA-binding domain |
| Pfam | PF00847 | 1.3E-9 | 322 | 378 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 17.056 | 323 | 386 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 5.2E-12 | 323 | 387 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 6.1E-22 | 323 | 392 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 3.9E-6 | 324 | 335 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 6.43E-25 | 421 | 482 | No hit | No description |
| SuperFamily | SSF54171 | 1.31E-17 | 421 | 481 | IPR016177 | DNA-binding domain |
| Gene3D | G3DSA:3.30.730.10 | 2.0E-18 | 422 | 480 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 8.3E-34 | 422 | 486 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 19.282 | 422 | 480 | IPR001471 | AP2/ERF domain |
| Pfam | PF00847 | 1.1E-9 | 423 | 472 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 3.9E-6 | 462 | 482 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007276 | Biological Process | gamete generation | ||||
| GO:0010492 | Biological Process | maintenance of shoot apical meristem identity | ||||
| GO:0042127 | Biological Process | regulation of cell proliferation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 661 aa Download sequence Send to blast |
MKSINNDDDN NSNTHWLGFS LSPLQMKMEI PSTSAASVCA SASSISVPTA IPASFFQSNI 60 NYGSYCAVEG ENGGLYSSLP VMPLKSDGSL CIMEALTRSQ TQATLAATTS TPKLEDFFGG 120 ATMGTHHYET TDREAMPLSL DSMYYNHNEP NHQNCLSHLH QNPRHQNHQI QVQHYPYYNN 180 NTNFRSYDML IGEEPKENQV SDSNVQIQNL GDDTLSGMRN WVSRNYSNNQ HAMEQQKMIS 240 CMGDSSGGES GAISAMAYGD LQSLSLSMSP GSQSSCVTGS QQISPAVTDC VAIETKKRGP 300 DKAEQKQTVH RKSIDTFGQR TSQYRGVTRH RWTGRYEAHL WDNSCKKEGQ SRKGRQGGYD 360 MEEKAARAYD LAALKYWGPS THINFQLENY QKELEEMKNM TRQEYVAHLR RKSSGFSRGA 420 SMYRGVTRHH QHGRWQARIG RVAGNKDLYL GTFSTQEEAA EAYDVAAIKF RGVNAVTNFD 480 ITRYDVERIM ASNTLVSGEV ARRNKDAGPV NEATNHSTHN SFSEVIPVQK NNQSESDWKM 540 ALYQSDQKAS NVIDNFKTQI FENVTGIDTL SSIHNQQVEE SGKIGTHLSN ASSLVTSLSS 600 SREGSPDRAS MPMLFAMPPS ATSKLFSNPS SNEISWIPTA AQLRSAVSLP HMPVFAAWTD 660 A |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00088 | SELEX | Transfer from AT4G37750 | Download |
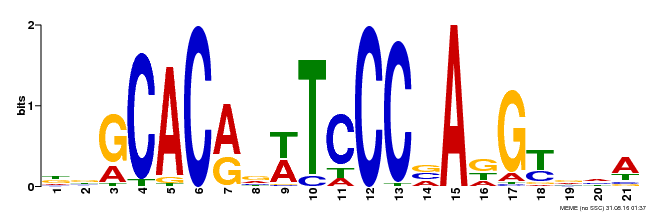
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_012079449.1 | 0.0 | AP2-like ethylene-responsive transcription factor ANT | ||||
| TrEMBL | A0A067KAR7 | 0.0 | A0A067KAR7_JATCU; Uncharacterized protein | ||||
| STRING | cassava4.1_003308m | 0.0 | (Manihot esculenta) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF916 | 34 | 114 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G37750.1 | 1e-147 | AP2 family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Entrez Gene | 105639879 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



