 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Itr_sc001678.1_g00001.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Convolvulaceae; Ipomoeeae; Ipomoea
|
||||||||
| Family | MIKC_MADS | ||||||||
| Protein Properties | Length: 278aa MW: 31488 Da PI: 6.544 | ||||||||
| Description | MIKC_MADS family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SRF-TF | 86.3 | 1.8e-27 | 9 | 59 | 1 | 51 |
S---SHHHHHHHHHHHHHHHHHHHHHHHHHHT-EEEEEEE-TTSEEEEEE- CS
SRF-TF 1 krienksnrqvtfskRrngilKKAeELSvLCdaevaviifsstgklyeyss 51
k ien nrqvtfskRr g+ KKA ELS+LCdaevavi+fsstgkl+e+ss
Itr_sc001678.1_g00001.1 9 KLIENINNRQVTFSKRRGGLVKKARELSILCDAEVAVIVFSSTGKLFEFSS 59
569**********************************************96 PP
| |||||||
| 2 | K-box | 57.1 | 7.7e-20 | 84 | 176 | 8 | 99 |
K-box 8 s.leeakaeslqqelakLkkeienLqreqRhllGedLesLslkeLqqLeqqLekslkkiRskKnellleqieelqkkekelqeenk 92
+e++ + + e++ L +ei++L+++q +lG+dL+ + l+eL++Le++L+++l i+s+K+++l+eq+e++qk+e+ + n
Itr_sc001678.1_g00001.1 84 PlVEKKPKQLEHMEVEVLSDEISKLKSKQLMFLGKDLKGMGLNELRELEHELHEGLLGIKSRKEQILMEQLEHSQKQEELALLVND 169
4456666777789************************************************************************* PP
K-box 93 aLrkkle 99
+Lr+++e
Itr_sc001678.1_g00001.1 170 QLRREIE 176
****986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00432 | 9.0E-40 | 1 | 60 | IPR002100 | Transcription factor, MADS-box |
| PROSITE profile | PS50066 | 31.23 | 1 | 61 | IPR002100 | Transcription factor, MADS-box |
| CDD | cd00265 | 8.10E-38 | 2 | 81 | No hit | No description |
| SuperFamily | SSF55455 | 1.96E-29 | 2 | 79 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 6.0E-28 | 3 | 23 | IPR002100 | Transcription factor, MADS-box |
| PROSITE pattern | PS00350 | 0 | 3 | 57 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF00319 | 1.4E-25 | 11 | 57 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 6.0E-28 | 23 | 38 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 6.0E-28 | 38 | 59 | IPR002100 | Transcription factor, MADS-box |
| PROSITE profile | PS51297 | 10.821 | 91 | 181 | IPR002487 | Transcription factor, K-box |
| Pfam | PF01486 | 6.6E-14 | 92 | 175 | IPR002487 | Transcription factor, K-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010047 | Biological Process | fruit dehiscence | ||||
| GO:0010262 | Biological Process | somatic embryogenesis | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0048577 | Biological Process | negative regulation of short-day photoperiodism, flowering | ||||
| GO:0060862 | Biological Process | negative regulation of floral organ abscission | ||||
| GO:0060867 | Biological Process | fruit abscission | ||||
| GO:0071365 | Biological Process | cellular response to auxin stimulus | ||||
| GO:2000692 | Biological Process | negative regulation of seed maturation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005737 | Cellular Component | cytoplasm | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 278 aa Download sequence Send to blast |
MGRGKIDIKL IENINNRQVT FSKRRGGLVK KARELSILCD AEVAVIVFSS TGKLFEFSST 60 RFAGNCMEQT LSRYNKRIKS SGMPLVEKKP KQLEHMEVEV LSDEISKLKS KQLMFLGKDL 120 KGMGLNELRE LEHELHEGLL GIKSRKEQIL MEQLEHSQKQ EELALLVNDQ LRREIEGFRG 180 FYYPLSATVP APSIEYCPTE NNVSSGKDGV ESQDIGSDGG FENENSDTTL QLGLAMGGYR 240 KRKTPEPETR STSLKLARGN EVDLAEQFIM KIQEWEEK |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5f28_A | 1e-18 | 1 | 74 | 1 | 69 | MEF2C |
| 5f28_B | 1e-18 | 1 | 74 | 1 | 69 | MEF2C |
| 5f28_C | 1e-18 | 1 | 74 | 1 | 69 | MEF2C |
| 5f28_D | 1e-18 | 1 | 74 | 1 | 69 | MEF2C |
| Search in ModeBase | ||||||
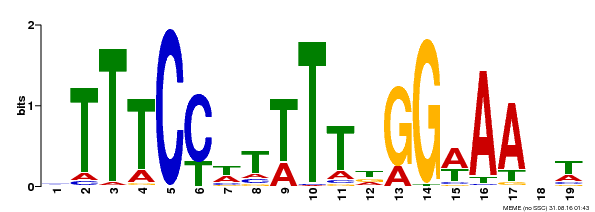
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00508 | DAP | Transfer from AT5G13790 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_019157010.1 | 1e-147 | PREDICTED: MADS-box transcription factor 23-like isoform X2 | ||||
| TrEMBL | A0A2U8UAC7 | 1e-103 | A0A2U8UAC7_IPOBA; MBP5 | ||||
| STRING | XP_009592797.1 | 1e-90 | (Nicotiana tomentosiformis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA8775 | 22 | 26 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G13790.1 | 6e-54 | AGAMOUS-like 15 | ||||



