 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Itr_sc000529.1_g00021.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Convolvulaceae; Ipomoeeae; Ipomoea
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 879aa MW: 100522 Da PI: 6.3171 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 83.2 | 4.4e-26 | 87 | 189 | 2 | 91 |
FAR1 2 fYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkk.............tekerrtraetrtgCkaklkvkkekdg 74
+Y+eYA+++GF++ +++s++sk+++e+++++f Cs++g+++e +k +e++ +ra +t+Cka+++vk++ dg
Itr_sc000529.1_g00021.1 87 YYQEYARSMGFNTAIQNSRRSKTSREFIDAKFACSRYGTKREYEKPanrprsrqgskqdQENATGRRACAKTDCKASMHVKRRPDG 172
8***************************************9999889999*9999886666666699******************* PP
FAR1 75 kwevtkleleHnHelap 91
kw ++++e+eHnHel+p
Itr_sc000529.1_g00021.1 173 KWIIHRFEKEHNHELMP 189
**************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 4.0E-24 | 87 | 189 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 1.0E-26 | 287 | 378 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 9.433 | 567 | 603 | IPR007527 | Zinc finger, SWIM-type |
| Pfam | PF04434 | 3.3E-5 | 567 | 600 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 4.2E-7 | 578 | 605 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009585 | Biological Process | red, far-red light phototransduction | ||||
| GO:0010218 | Biological Process | response to far red light | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 879 aa Download sequence Send to blast |
MDIDLRLPSG DHDKEEETNG IVNLLDHEEK LDGVEVGGDM DIEEKIHVED GGEMNSPMGD 60 MGEFKEDVNL EPLAGMEFES HGEAYAYYQE YARSMGFNTA IQNSRRSKTS REFIDAKFAC 120 SRYGTKREYE KPANRPRSRQ GSKQDQENAT GRRACAKTDC KASMHVKRRP DGKWIIHRFE 180 KEHNHELMPA QAVSEQTRRM YAAMARQFAE YKNVVGLKHD SRSPSDKVRN LAMEAGEANI 240 LLEFFIQMQS MNSNFFYAID VSEDHRLKNL FWTDAKSRHD YANFSDVVSF DTTYLRNKYK 300 MPLALFIGVN QHYQFMLLGC ALLSDESSAT FSWVMRTWLK AMGDQAPKII ITDQEMVMKS 360 VVSEILPSTL HFFCQWQVLG KVSENLNHII KQNEEFMAEF ENCIYRSWTD EEFEKSWQNL 420 VQKFELGENE LMLSLYEDRT KWVPTFMKNA FLAGMSTALR SESVNSFFDK YVHRKTTIQE 480 FVKQYEAILQ DRYEEEAKAD SDTWNKQPAL KSPSPFEKHV AGVYTHAVFK KFQVEVVAAV 540 ACIPKDEKRD ETAVTFIIQD FEKSQEFTVT WNGHKSEVSC MCHMFEYKGF LCRHAMVVLQ 600 IGGISTIPPQ YILKRWTKDA KVRFPVIDGS EGVQSRVQRY NSLSQQAMRL AEEGSLSQES 660 YRFAVRSLDE TFGNCVDANS SNKNLVDAGT TSTPGLLCTE EDNQNRSMSK MNKKKNNPTK 720 KRKANSEPDV MAVGAPESLQ QMDKLSSRPV TLDGYFGPQH GLQGMVQLNL MAPTRDNYYA 780 NQQPMQGLGQ LNSIAPTHDG YYGTQPAMHG LGQMDFFRTP SYPYGIRVNI TAFSQILENQ 840 LIKCLSESSF VSSCAGAHGS VEVKIFYNAI WFVRRALDL |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
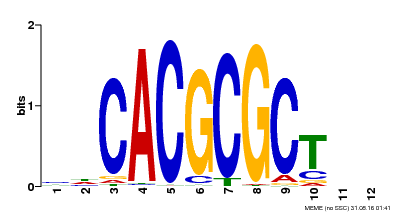
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. When associated with PHYA, protects it from being recognized and degraded by the COP1/SPA complex. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:17012604, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:18715961, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00078 | ChIP-seq | Transfer from AT3G22170 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. Up-regulated by white light. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_019153175.1 | 0.0 | PREDICTED: protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| Refseq | XP_019153177.1 | 0.0 | PREDICTED: protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| Refseq | XP_019153178.1 | 0.0 | PREDICTED: protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| Refseq | XP_019153179.1 | 0.0 | PREDICTED: protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| Swissprot | Q9LIE5 | 0.0 | FHY3_ARATH; Protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| TrEMBL | A0A1S4AZK2 | 0.0 | A0A1S4AZK2_TOBAC; protein FAR-RED ELONGATED HYPOCOTYL 3-like isoform X3 | ||||
| TrEMBL | A0A1U7YPD6 | 0.0 | A0A1U7YPD6_NICSY; protein FAR-RED ELONGATED HYPOCOTYL 3 isoform X3 | ||||
| STRING | XP_009804946.1 | 0.0 | (Nicotiana sylvestris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA496 | 21 | 103 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G22170.2 | 0.0 | far-red elongated hypocotyls 3 | ||||



