 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Itr_sc000111.1_g00057.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Convolvulaceae; Ipomoeeae; Ipomoea
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 295aa MW: 32516.6 Da PI: 6.5151 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 42.7 | 1.3e-13 | 38 | 102 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT...................-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGgg...................tWktIartmgkgRtlkqcksrwqky 47
r +WT+eE++++++a ++l ++ +Wk+I +g +t q++s+ qky
Itr_sc000111.1_g00057.1 38 RESWTEEEHDKFLEALQLLSTPlppssiyifwpyslflfdrDWKKIEDFVG-SKTVIQIRSHAQKY 102
789************************************************.*************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 3.34E-13 | 32 | 108 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50090 | 6.005 | 33 | 103 | IPR017877 | Myb-like domain |
| Gene3D | G3DSA:1.10.10.60 | 1.0E-6 | 36 | 108 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 1.0E-6 | 37 | 105 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 7.1E-10 | 38 | 102 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 3.11E-6 | 40 | 103 | No hit | No description |
| TIGRFAMs | TIGR01557 | 1.7E-7 | 76 | 105 | IPR006447 | Myb domain, plants |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0032922 | Biological Process | circadian regulation of gene expression | ||||
| GO:0043966 | Biological Process | histone H3 acetylation | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0048573 | Biological Process | photoperiodism, flowering | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 295 aa Download sequence Send to blast |
MTSASSNSQS MANPSNSSTP TDGSGKKVRK PYTITKSRES WTEEEHDKFL EALQLLSTPL 60 PPSSIYIFWP YSLFLFDRDW KKIEDFVGSK TVIQIRSHAQ KYFLKVQKNG TIAHVPPPRP 120 KRKASHPYPQ KAPKNVLLPL PGSMAYPSSV NPLAPGYPSW DETSMLVNAS TSGMMPPEDE 180 FNVHGVEADV GSKGAARIGN SDVGEIGSSN RTVPNQSKLG SVLHGLPDFA EVYSFIGSVF 240 DPDTSFHELK LKEMDPINLE TVSVGTDETS SYLFNDYAYY VTGGWKCVHI SCEIS |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator of evening element (EE)-containing clock-controlled genes. Forms a negative feedback loop with APRR5. Regulates the pattern of histone H3 acetylation of the TOC1 promoter. {ECO:0000269|PubMed:21205033, ECO:0000269|PubMed:21474993, ECO:0000269|PubMed:21483796, ECO:0000269|PubMed:23638299}. | |||||
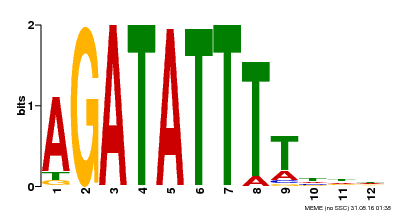
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00338 | DAP | Transfer from AT3G09600 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Circadian-regulation. Peak of expression in the afternoon. Down-regulated by cold. {ECO:0000269|PubMed:21205033, ECO:0000269|PubMed:22902701, ECO:0000269|PubMed:23638299}. | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AM455206 | 6e-32 | AM455206.2 Vitis vinifera contig VV78X069637.42, whole genome shotgun sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_019184918.1 | 1e-164 | PREDICTED: protein REVEILLE 8 isoform X1 | ||||
| Swissprot | Q8RWU3 | 2e-99 | RVE8_ARATH; Protein REVEILLE 8 | ||||
| TrEMBL | A0A2G3BG26 | 1e-125 | A0A2G3BG26_CAPCH; Protein REVEILLE 6 | ||||
| STRING | Solyc10g084370.1.1 | 1e-123 | (Solanum lycopersicum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA998 | 24 | 78 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G09600.1 | 1e-102 | MYB_related family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



