 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MLOC_9943.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Hordeinae; Hordeum
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 272aa MW: 29999.7 Da PI: 6.8033 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 41.5 | 3.1e-13 | 223 | 270 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHT...TTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmg...kgRtlkqcksrwqky 47
rW+ E+e l+d+v+q+G+g+Wk I ++ gRt+ ++k++w++
MLOC_9943.1 223 RWSSVEEEALKDGVEQFGSGNWKDILSHNAdvfIGRTPVDLKDKWRNM 270
9********************************9************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 15.904 | 216 | 272 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.5E-14 | 216 | 272 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 1.87E-15 | 217 | 271 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 3.0E-5 | 220 | 272 | IPR001005 | SANT/Myb domain |
| CDD | cd11660 | 5.37E-20 | 222 | 272 | No hit | No description |
| Pfam | PF00249 | 4.2E-10 | 223 | 270 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009901 | Biological Process | anther dehiscence | ||||
| GO:0010152 | Biological Process | pollen maturation | ||||
| GO:0043067 | Biological Process | regulation of programmed cell death | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 272 aa Download sequence Send to blast |
GNAFSQEALK GLFGDRAKAE RRVRDLLAAE WAAIGPSRLE LAAEQIAGDG AVETWRAADE 60 TVRAKYRILV GEEKAREILS RIEDPISSPQ VHKVIHDLKS SCADLHNVVD DPLPAAKAAA 120 DKVLAARMDN AVNINAEVNN QAANCSIAGS SAVNDQGETL RKGMPSSLMD WNPTAQSLLW 180 EDSIDSDGSR SQLHRPHLPS PRRIPVSPLQ VAENKSMRRR ARRWSSVEEE ALKDGVEQFG 240 SGNWKDILSH NADVFIGRTP VDLKDKWRNM MR |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00052 | PBM | Transfer from AT1G15720 | Download |
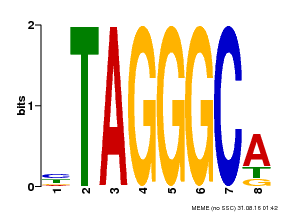
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | MLOC_9943.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK375507 | 0.0 | AK375507.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, complete cds, clone: NIASHv3096E16. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020185046.1 | 1e-168 | uncharacterized protein LOC109770747 | ||||
| TrEMBL | F2EH80 | 0.0 | F2EH80_HORVV; Predicted protein | ||||
| STRING | MLOC_9943.1 | 0.0 | (Hordeum vulgare) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP3154 | 30 | 40 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G15720.1 | 4e-15 | TRF-like 5 | ||||



