 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MLOC_68907.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Hordeinae; Hordeum
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 747aa MW: 83941.8 Da PI: 6.725 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 72.3 | 6e-23 | 72 | 173 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT.......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSEE CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrsef 90
f+k+lt sd++++g +++ +++a+e+ +++++ ++l+ +d++g W++++i+r++++r++l++GW+ Fv++++L +gD ++F + +++
MLOC_68907.1 72 FCKTLTASDTSTHGGFSVLRRHADEClpsldmtQSPPT--QELVAKDLHGMDWRFRHIFRGQPRRHLLQSGWSVFVSSKRLVAGDAFIFL--RGESG 164
99****************************97555544..58************************************************..44999 PP
..EEEEE-S CS
B3 91 elvvkvfrk 99
el+v+v+r+
MLOC_68907.1 165 ELRVGVRRA 173
9*****996 PP
| |||||||
| 2 | Auxin_resp | 111.4 | 9.1e-37 | 198 | 279 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrlsGtvvgvsdldpvrWpnSkWrsLk 83
a+ha++tks+F+v+Y+Pr+s+seF+++++++++++k+++s+GmRf+m+fe+e+++e+r++Gt+vg ++ld+ Wp+S+WrsLk
MLOC_68907.1 198 AWHAINTKSMFTVYYKPRTSPSEFIIPYDQYMESVKNNYSIGMRFRMRFEGEEAPEQRFTGTIVGSENLDQ-LWPESNWRSLK 279
89******************************************************************999.9********96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101936 | 6.41E-40 | 65 | 201 | IPR015300 | DNA-binding pseudobarrel domain |
| Gene3D | G3DSA:2.40.330.10 | 3.8E-39 | 65 | 187 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 3.63E-19 | 70 | 172 | No hit | No description |
| Pfam | PF02362 | 8.6E-21 | 72 | 173 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 4.5E-21 | 72 | 174 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 11.702 | 72 | 174 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 1.1E-33 | 198 | 279 | IPR010525 | Auxin response factor |
| Pfam | PF02309 | 3.4E-5 | 608 | 662 | IPR033389 | AUX/IAA domain |
| SuperFamily | SSF54277 | 1.13E-12 | 620 | 703 | No hit | No description |
| PROSITE profile | PS51745 | 25.01 | 626 | 719 | IPR000270 | PB1 domain |
| Pfam | PF02309 | 9.5E-8 | 667 | 710 | IPR033389 | AUX/IAA domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008285 | Biological Process | negative regulation of cell proliferation | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009911 | Biological Process | positive regulation of flower development | ||||
| GO:0010047 | Biological Process | fruit dehiscence | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0010227 | Biological Process | floral organ abscission | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0048481 | Biological Process | plant ovule development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 747 aa Download sequence Send to blast |
MNQVAGNQMR LYDLPPKLLC RVINVELKAE ADTDEVYAQV MLMPEPEQNE MAVDKSTSTT 60 GATPPRPAVR SFCKTLTASD TSTHGGFSVL RRHADECLPS LDMTQSPPTQ ELVAKDLHGM 120 DWRFRHIFRG QPRRHLLQSG WSVFVSSKRL VAGDAFIFLR GESGELRVGV RRAMRQLSNV 180 PSSVISSHSM HLGVLATAWH AINTKSMFTV YYKPRTSPSE FIIPYDQYME SVKNNYSIGM 240 RFRMRFEGEE APEQRFTGTI VGSENLDQLW PESNWRSLKV RWDEPSTIPR PDRVSPWKIE 300 PASSPPVNPL PLSRVKRPRP NVPPVSPESS VLTKEGATKI DMDSAQAQQR NQNNMVLQGQ 360 EHMTLRTNNL TGSNDSDATV QKPMMWSPSP NIGKNHASAF QQRPSMENWM QLGRCETAFK 420 DASSGAQSFG DSQGFFMQTF DEAPNRHGSF KNQFQDHSSA RHFSDPYTKM HTEANEFHFW 480 NSQSTVYGNP RDQSQGFRFE EHPSNWLRQQ QFSPVEQPRV IRPQASIAPV DLEKAREGSG 540 FKIFGFKVDT TSAPSNHLSS TMAVIHEPVL QTQASASLTQ LQHAHIDCIP ELSVSTAGTT 600 ENEKSIQQAP NSSKDVQSKS HGASTRSCTK VHKQGVALGR SVDLSKFGDY DELTAELDRM 660 FEFDGELMSS NRDWQIVYTD PEGDMMLVGD DPWEEFCSIV RKIFIYTKEE VQKMNSKSST 720 PRKEEGSADA DGANEKAHLA ASSHLDN |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldv_A | 1e-142 | 1 | 304 | 56 | 357 | Auxin response factor 1 |
| 4ldw_A | 1e-142 | 1 | 304 | 56 | 357 | Auxin response factor 1 |
| 4ldw_B | 1e-142 | 1 | 304 | 56 | 357 | Auxin response factor 1 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 315 | 319 | KRPRP |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). | |||||
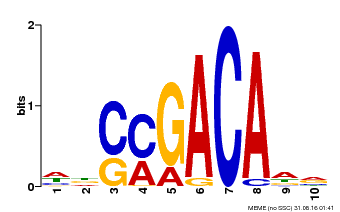
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00574 | DAP | Transfer from AT5G62000 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | MLOC_68907.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK364852 | 0.0 | AK364852.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, complete cds, clone: NIASHv2028N02. | |||
| GenBank | AK368785 | 0.0 | AK368785.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, complete cds, clone: NIASHv2079M07. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020198715.1 | 0.0 | auxin response factor 4-like | ||||
| Swissprot | Q5JK20 | 0.0 | ARFD_ORYSJ; Auxin response factor 4 | ||||
| TrEMBL | A0A287MEW0 | 0.0 | A0A287MEW0_HORVV; Auxin response factor | ||||
| STRING | MLOC_68907.1 | 0.0 | (Hordeum vulgare) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1178 | 36 | 111 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G62000.3 | 0.0 | auxin response factor 2 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



