 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MLOC_64817.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Hordeinae; Hordeum
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 424aa MW: 44704.7 Da PI: 9.864 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 55.4 | 1.4e-17 | 116 | 161 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+g+W++eEd+ l ++v ++G ++W++I++ ++ gR++k+c++rw +
MLOC_64817.1 116 KGPWSPEEDDALQRLVGRHGARNWSLISKSIP-GRSGKSCRLRWCNQ 161
79******************************.***********985 PP
| |||||||
| 2 | Myb_DNA-binding | 51.4 | 2.6e-16 | 170 | 212 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
++T++Ed+ +++a++++G++ W+tIar + gRt++ +k++w++
MLOC_64817.1 170 PFTPDEDDTILRAHARFGNK-WATIARLLS-GRTDNAIKNHWNST 212
89******************.*********.***********986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 25.716 | 111 | 166 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 4.87E-31 | 113 | 209 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 1.0E-15 | 115 | 164 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.0E-17 | 116 | 161 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 6.6E-25 | 117 | 169 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 1.16E-14 | 118 | 160 | No hit | No description |
| SMART | SM00717 | 7.2E-14 | 167 | 215 | IPR001005 | SANT/Myb domain |
| PROSITE profile | PS51294 | 19.747 | 168 | 217 | IPR017930 | Myb domain |
| CDD | cd00167 | 7.23E-11 | 170 | 213 | No hit | No description |
| Pfam | PF00249 | 7.8E-14 | 170 | 212 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 9.0E-23 | 170 | 216 | IPR009057 | Homeodomain-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 424 aa Download sequence Send to blast |
MVATPRVPGP AVPTSPPSLP CWAGTRSPSP PVPGIYPPPA PSAPLTPSFL CLLPSSLHFS 60 TAPPPPHYFG NPPPVAGSGY YAHVHPSIDR SIYGRRVMAA CRGGGGGGGK DVDRIKGPWS 120 PEEDDALQRL VGRHGARNWS LISKSIPGRS GKSCRLRWCN QLSPQVEHRP FTPDEDDTIL 180 RAHARFGNKW ATIARLLSGR TDNAIKNHWN STLKRKYSSS SSASASASAS ASPGDDAVDD 240 DRRPLKRTSS DGYPGLCFSP GSPSGSDLSD SSHQSLPSVM PSAASHQPHV YRPVARAGGV 300 VVLPTATPQP SPPSPPPPPP PPPPATSLSL SLSLPGLDAP EPAPVAQPMP MPMPPPPVPA 360 PVQPAAAPQM PPPSMPFHLQ PPPPPQPRAA APFNGEFLSM MQEMIRIEVR NYMSGFHPRS 420 PTSA |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 1e-38 | 113 | 216 | 4 | 107 | B-MYB |
| 1gv2_A | 9e-39 | 115 | 216 | 3 | 104 | MYB PROTO-ONCOGENE PROTEIN |
| 1mse_C | 9e-39 | 115 | 216 | 3 | 104 | C-Myb DNA-Binding Domain |
| 1msf_C | 9e-39 | 115 | 216 | 3 | 104 | C-Myb DNA-Binding Domain |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 101 | 109 | RGGGGGGGK |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00476 | DAP | Transfer from AT4G37260 | Download |
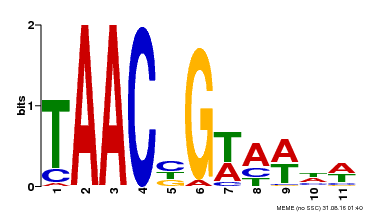
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | MLOC_64817.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK355241 | 0.0 | AK355241.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, complete cds, clone: NIASHv1018A13. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020146152.1 | 1e-159 | transcription factor MYB44-like | ||||
| TrEMBL | M0Y130 | 0.0 | M0Y130_HORVV; Uncharacterized protein | ||||
| STRING | MLOC_64817.1 | 0.0 | (Hordeum vulgare) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP789 | 38 | 154 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G23290.1 | 2e-68 | myb domain protein 70 | ||||



