 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MLOC_64597.3 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Hordeinae; Hordeum
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 479aa MW: 52803 Da PI: 10.0925 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 97.9 | 7.2e-31 | 110 | 163 | 2 | 55 |
G2-like 2 prlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
prlrWtp+LH +F++ave+LGG+e+AtPk +l++m+v gL+++hvkSHLQ+YR+
MLOC_64597.3 110 PRLRWTPDLHMAFIRAVERLGGQERATPKLVLQMMNVRGLSIAHVKSHLQMYRS 163
8****************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 12.702 | 106 | 166 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 3.6E-28 | 106 | 164 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 8.6E-14 | 109 | 164 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 4.1E-22 | 110 | 165 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 5.1E-9 | 111 | 162 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 479 aa Download sequence Send to blast |
MITREKEAIV RDVEDKQEVA EPEIVMDNRK RTMNGRKLDL NEDIDMESEQ GEVGDDNDRK 60 EEEEDGGSTT DVAGSRSSSN NSSTNHASET QEGTGAGEHR VRQYNRSKLP RLRWTPDLHM 120 AFIRAVERLG GQERATPKLV LQMMNVRGLS IAHVKSHLQM YRSKKLDHQG QRIRGAISSV 180 FSPMGFHSMR GDRRFHDMLL QRAAALSSTA EHGGFFASRN GGGGGNTTSR LYGILQHHHR 240 PSLIQSGFKN CSFRNQEWAF NHRDRITRKD VKASSTTPHF FASSSVRRWP LGSAVAGAGE 300 QMRDESFGYF TGHGSGQHSR AMPPARSVVP GGGNHRLPFR WHGSDGGKGA QATPSDPVMI 360 AKALDSQQQK HIEQQRALIT PADKVRLPTE TPELELSLSP ATAMDTTDGT PSKKRKKSTA 420 ASSEQELEHC NKLSISPLSL LPPAASMPMQ RKEKTRGGSA EAALGQSTLD LTMSIRALE |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 5e-18 | 111 | 163 | 4 | 56 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 5e-18 | 111 | 163 | 4 | 56 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_A | 4e-18 | 111 | 163 | 3 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 4e-18 | 111 | 163 | 3 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 4e-18 | 111 | 163 | 3 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 4e-18 | 111 | 163 | 3 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 5e-18 | 111 | 163 | 4 | 56 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 5e-18 | 111 | 163 | 4 | 56 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 5e-18 | 111 | 163 | 4 | 56 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 5e-18 | 111 | 163 | 4 | 56 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 5e-18 | 111 | 163 | 4 | 56 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 5e-18 | 111 | 163 | 4 | 56 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 412 | 416 | KKRKK |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00297 | DAP | Transfer from AT2G38300 | Download |
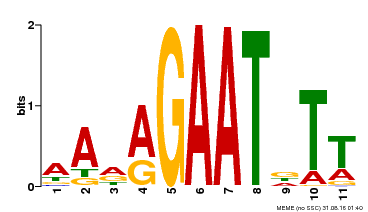
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | MLOC_64597.3 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020187256.1 | 0.0 | uncharacterized protein LOC109772958 isoform X1 | ||||
| TrEMBL | M0Y082 | 0.0 | M0Y082_HORVV; Uncharacterized protein | ||||
| STRING | MLOC_64597.2 | 0.0 | (Hordeum vulgare) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G38300.1 | 3e-37 | G2-like family protein | ||||



