 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MLOC_56567.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Hordeinae; Hordeum
|
||||||||
| Family | RAV | ||||||||
| Protein Properties | Length: 348aa MW: 38249.4 Da PI: 10.1454 | ||||||||
| Description | RAV family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 34.5 | 5.3e-11 | 55 | 103 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkr.krfslgkfgtaeeAakaaiaarkkleg 55
s++kGV + +grW A+I++ r +r++lg+f + Aa+a++ a ++++g
MLOC_56567.1 55 SRFKGVVPQP-NGRWGAQIYE------RhARVWLGTFPDQDSAARAYDVASLRYRG 103
689***9788.8*********......44*************************98 PP
| |||||||
| 2 | B3 | 93.1 | 1.9e-29 | 169 | 264 | 1 | 87 |
EEEE-..-HHHHTT-EE--HHH.HTT.........---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SS CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.........ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgr 87
f+k+ tpsdv+k++rlv+pk++ae+h + +++ l +ed +g++W+++++y+++s++yvltkGW++Fv+++gL +gD+++F+ +
MLOC_56567.1 169 FEKAVTPSDVGKLNRLVVPKQHAEKHfplkrspetTTTTGNGVLLNFEDGQGKVWRFRYSYWNSSQSYVLTKGWSRFVREKGLGAGDSIMFSCSAY 264
89*************************99877666444448999***********************************************96543 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF54171 | 2.03E-15 | 55 | 112 | IPR016177 | DNA-binding domain |
| CDD | cd00018 | 9.99E-21 | 55 | 111 | No hit | No description |
| Pfam | PF00847 | 8.5E-7 | 55 | 103 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 2.1E-18 | 56 | 111 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 20.192 | 56 | 111 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 3.6E-21 | 56 | 117 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:2.40.330.10 | 4.0E-37 | 165 | 279 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 3.79E-29 | 166 | 274 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 3.80E-28 | 167 | 259 | No hit | No description |
| SMART | SM01019 | 9.7E-22 | 169 | 277 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 13.296 | 169 | 278 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 5.1E-27 | 169 | 264 | IPR003340 | B3 DNA binding domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 348 aa Download sequence Send to blast |
MGVEILSSMV EHSFQYSSGA SSATAESGAV GTPPRHLSLP VAIADESLTS RSASSRFKGV 60 VPQPNGRWGA QIYERHARVW LGTFPDQDSA ARAYDVASLR YRGGDAAFNF PCVVVEAELA 120 FLAAHSKAEI VDMLRKQTYA DELRQGLRRG RGMGVRAQPM PSWARVPLFE KAVTPSDVGK 180 LNRLVVPKQH AEKHFPLKRS PETTTTTGNG VLLNFEDGQG KVWRFRYSYW NSSQSYVLTK 240 GWSRFVREKG LGAGDSIMFS CSAYGQEKQF FIDCKKNTTV NGGKSASPLQ VMEIAKAEQV 300 RVVRLFGVDI AGVKRERAAT AEQGPQGWFK RQCMAHGQHS PALGDFAL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1wid_A | 5e-46 | 168 | 275 | 13 | 115 | DNA-binding protein RAV1 |
| Search in ModeBase | ||||||
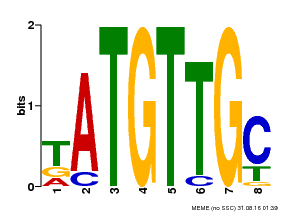
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00025 | PBM | Transfer from AT1G68840 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | MLOC_56567.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK362038 | 0.0 | AK362038.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, complete cds, clone: NIASHv2001P16. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020191173.1 | 0.0 | AP2/ERF and B3 domain-containing protein Os01g0141000-like isoform X1 | ||||
| Refseq | XP_020200371.1 | 0.0 | AP2/ERF and B3 domain-containing protein Os01g0141000-like isoform X1 | ||||
| Refseq | XP_020201146.1 | 0.0 | AP2/ERF and B3 domain-containing protein Os01g0141000-like isoform X1 | ||||
| Swissprot | Q9AWS0 | 1e-132 | Y1410_ORYSJ; AP2/ERF and B3 domain-containing protein Os01g0141000 | ||||
| TrEMBL | F2DDS1 | 0.0 | F2DDS1_HORVV; Predicted protein | ||||
| STRING | MLOC_56567.1 | 0.0 | (Hordeum vulgare) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP456 | 38 | 203 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G68840.2 | 2e-94 | related to ABI3/VP1 2 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



