 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MLOC_52321.2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Hordeinae; Hordeum
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 360aa MW: 38687.8 Da PI: 8.9463 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 132.9 | 1.1e-41 | 28 | 104 | 2 | 78 |
-SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 2 CqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
C vegC+adls+++eyhrrhkvCe+hsk+pvv+v+g++qrfCqqCsrfh l efDe krsCr+rL++hn+rrrk+q+
MLOC_52321.2 28 CSVEGCTADLSRCREYHRRHKVCEAHSKTPVVAVAGQQQRFCQQCSRFHLLGEFDEVKRSCRKRLDGHNRRRRKPQP 104
**************************************************************************987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 2.2E-34 | 19 | 89 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.137 | 25 | 102 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 2.49E-39 | 26 | 106 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 3.5E-30 | 28 | 101 | IPR004333 | Transcription factor, SBP-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0048653 | Biological Process | anther development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 360 aa Download sequence Send to blast |
QQQQQQPPSA ATAKRARAGQ GQQTVPACSV EGCTADLSRC REYHRRHKVC EAHSKTPVVA 60 VAGQQQRFCQ QCSRFHLLGE FDEVKRSCRK RLDGHNRRRR KPQPDPLNPA GLFANHHGVP 120 RFASYPQIFS PTSMAEPKWP GGIAVKTEAD AFHEQYYSFN GAASLFHHGK PERKHFPFLT 180 DGGDATFGCQ PPAFTITPSS ESSSNSSRHS NGKTTMFAHD GGPDHNCALS LLSDNPAPAH 240 IMNPAEAHHL GGGGGVTIQY GGGGGGKVAR LHNNGDVSLT GLSYVSLGDK GAPMLLHTSN 300 RSQHAAATAT TAVTTSTAPA PTASQMQQYH GYYHHQHQVG ADQGNHQDAG GMHALPFSSW |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 9e-28 | 21 | 101 | 4 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 84 | 101 | KRSCRKRLDGHNRRRRKP |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' (By similarity). May be involved in panicle development. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00555 | DAP | Transfer from AT5G50570 | Download |
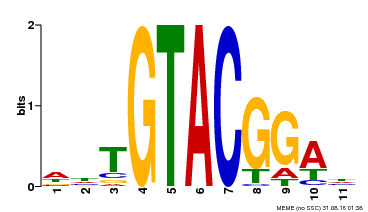
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | MLOC_52321.2 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Negatively regulated by microRNAs miR156b and miR156h. {ECO:0000305|PubMed:16861571}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK356077 | 0.0 | AK356077.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, complete cds, clone: NIASHv1029L02. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020178580.1 | 0.0 | squamosa promoter-binding-like protein 2 | ||||
| Swissprot | Q0JGI1 | 1e-108 | SPL2_ORYSJ; Squamosa promoter-binding-like protein 2 | ||||
| TrEMBL | F2CWS4 | 0.0 | F2CWS4_HORVV; Predicted protein | ||||
| STRING | MLOC_52321.2 | 0.0 | (Hordeum vulgare) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP10456 | 30 | 38 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G50670.1 | 1e-35 | SBP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



