 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MLOC_3855.2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Hordeinae; Hordeum
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 265aa MW: 30427.5 Da PI: 7.996 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 17.7 | 9.7e-06 | 9 | 34 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
++Cp Cgk+Fs k n +rH + H
MLOC_3855.2 9 FTCPleGCGKRFSIKANIQRHVKEmH 34
89********************9988 PP
| |||||||
| 2 | zf-C2H2 | 17.3 | 1.3e-05 | 53 | 75 | 3 | 23 |
ET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 3 Cp..dCgksFsrksnLkrHirtH 23
C+ C+k F+ +s+Lk+H +H
MLOC_3855.2 53 CQeeGCKKAFKYPSQLKKHEEMH 75
77779**************9988 PP
| |||||||
| 3 | zf-C2H2 | 16.3 | 2.7e-05 | 142 | 166 | 2 | 23 |
EET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 2 kCp..dCgksFsrksnLkrHirt.H 23
kC+ C +Fs+ksnL++H++ H
MLOC_3855.2 142 KCTfdGCECTFSNKSNLNKHMKAcH 166
799999****************955 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:3.30.160.60 | 5.5E-7 | 2 | 31 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 7.59E-7 | 7 | 35 | No hit | No description |
| SMART | SM00355 | 4.0E-4 | 9 | 34 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 12.217 | 9 | 39 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 11 | 34 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.032 | 51 | 75 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 9.349 | 51 | 76 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 53 | 75 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 7.3E-7 | 53 | 75 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 3.5 | 83 | 108 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 85 | 108 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 19 | 111 | 132 | IPR015880 | Zinc finger, C2H2-like |
| SuperFamily | SSF57667 | 2.11E-9 | 126 | 183 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 7.0E-7 | 126 | 163 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0015 | 141 | 166 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.115 | 141 | 171 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 143 | 166 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 6.0E-4 | 165 | 194 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 27 | 172 | 198 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 8.517 | 172 | 198 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 174 | 198 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008097 | Molecular Function | 5S rRNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0080084 | Molecular Function | 5S rDNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 265 aa Download sequence Send to blast |
MLTHEGKLFT CPLEGCGKRF SIKANIQRHV KEMHEDEHEC KMDDAKSNQQ VICQEEGCKK 60 AFKYPSQLKK HEEMHAKLDY VEVVCCEPGC LKTFSNAGCL KAHNQSCHLY TQCDICGEKH 120 LKKNIKRHLR AHAEVHSTER LKCTFDGCEC TFSNKSNLNK HMKACHDQVR PFECRVVGCG 180 KTFHYKHVRD NHEKSSWHVY VEGDYEEMDE QLRSRPRGGR KRAAVTVETL TRKRVTIPSE 240 ASSLDDGAGY MRWLLSGGDD SNHAE |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Essential protein (PubMed:22353599). Isoform 1 is a transcription activator the binds both 5S rDNA and 5S rRNA and stimulates the transcription of 5S rRNA gene (PubMed:12711688, PubMed:22353599). Isoform 1 regulates 5S rRNA levels during development (PubMed:22353599). {ECO:0000269|PubMed:12711688, ECO:0000269|PubMed:22353599}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00229 | DAP | Transfer from AT1G72050 | Download |
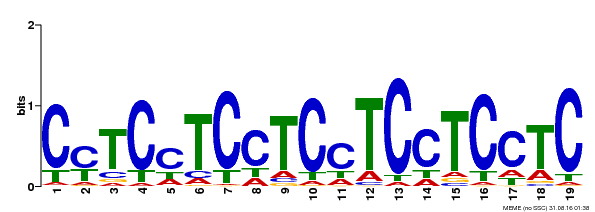
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | MLOC_3855.2 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK374527 | 0.0 | AK374527.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, complete cds, clone: NIASHv3068C12. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020181263.1 | 0.0 | transcription factor IIIA-like | ||||
| Swissprot | Q84MZ4 | 5e-86 | TF3A_ARATH; Transcription factor IIIA | ||||
| TrEMBL | M0VYG1 | 0.0 | M0VYG1_HORVV; Uncharacterized protein | ||||
| TrEMBL | M0VYG2 | 0.0 | M0VYG2_HORVV; Uncharacterized protein | ||||
| STRING | MLOC_3855.1 | 0.0 | (Hordeum vulgare) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G72050.1 | 2e-89 | transcription factor IIIA | ||||



