 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MLOC_34885.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Hordeinae; Hordeum
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 226aa MW: 24594.9 Da PI: 9.9795 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 54.6 | 2.6e-17 | 14 | 61 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT+eEd+ lv +++++G+g+W++++ g+ R+ k+c++rw +yl
MLOC_34885.1 14 KGPWTPEEDLMLVSYIQEHGPGNWRAVPTNTGLMRCSKSCRLRWTNYL 61
79********************************************97 PP
| |||||||
| 2 | Myb_DNA-binding | 48.3 | 2.3e-15 | 67 | 112 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rg++ +E++l+v++ ++lG++ W++Ia++++ Rt++++k++w+++l
MLOC_34885.1 67 RGNFNDQEEKLIVHLQALLGNR-WAAIASYLP-ERTDNDIKNYWNTHL 112
899*******************.*********.************996 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 3.6E-24 | 5 | 63 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 24.713 | 9 | 65 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 8.39E-30 | 10 | 108 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 1.3E-13 | 13 | 63 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.8E-16 | 14 | 61 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 7.91E-12 | 16 | 61 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 1.3E-23 | 64 | 117 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 19.345 | 66 | 116 | IPR017930 | Myb domain |
| SMART | SM00717 | 2.8E-15 | 66 | 114 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 4.7E-14 | 67 | 112 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 4.73E-11 | 69 | 112 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0080167 | Biological Process | response to karrikin | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 226 aa Download sequence Send to blast |
MGRPPCCDKM GVKKGPWTPE EDLMLVSYIQ EHGPGNWRAV PTNTGLMRCS KSCRLRWTNY 60 LRPGIKRGNF NDQEEKLIVH LQALLGNRWA AIASYLPERT DNDIKNYWNT HLKKKLKKMQ 120 DAGGGGHDGG SEGAGVTKAA APKGQWERRL QTDIHTARQA LRDALSLEPS QPTAMAVPAL 180 PTPPGSVTTY ASSADNIARL LEGWMRPGSS SKGPEASGST SSTTAT |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1gv2_A | 7e-26 | 14 | 116 | 4 | 105 | MYB PROTO-ONCOGENE PROTEIN |
| 1h8a_C | 1e-25 | 12 | 116 | 25 | 128 | MYB TRANSFORMING PROTEIN |
| 1mse_C | 7e-26 | 14 | 116 | 4 | 105 | C-Myb DNA-Binding Domain |
| 1msf_C | 7e-26 | 14 | 116 | 4 | 105 | C-Myb DNA-Binding Domain |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator involved in the activation of cuticular wax biosynthesis under drought stress. Binds directly to the promoters of genes involved in cuticular wax biosynthesis. Transactivates WSD1, KCS2/DAISY, CER1, CER2, FAR3 and ECR genes (PubMed:25305760, PubMed:27577115). Functions together with MYB96 in the activation of cuticular wax biosynthesis (PubMed:27577115). {ECO:0000269|PubMed:25305760, ECO:0000269|PubMed:27577115}. | |||||
| UniProt | Transcription factor. {ECO:0000305}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00395 | DAP | Transfer from AT3G47600 | Download |
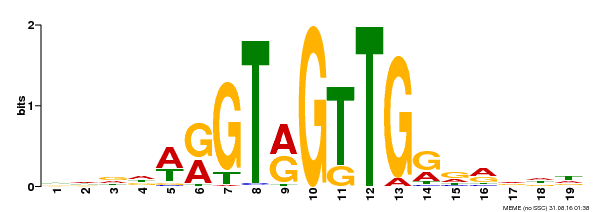
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | MLOC_34885.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by salt stress, osmotic shock and abscisic acid (ABA). {ECO:0000269|PubMed:25305760}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | JF951914 | 0.0 | JF951914.1 Aegilops tauschii clone TaMYB31 R2R3-MYB protein mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020146652.1 | 1e-133 | myb-related protein 306-like | ||||
| Swissprot | P81392 | 1e-104 | MYB06_ANTMA; Myb-related protein 306 | ||||
| Swissprot | Q9SN78 | 1e-104 | MYB94_ARATH; Transcription factor MYB94 | ||||
| TrEMBL | A0A3B6MSP3 | 1e-131 | A0A3B6MSP3_WHEAT; MYB transcription factor 31-D | ||||
| TrEMBL | G9DR77 | 1e-131 | G9DR77_AEGTA; MYB transcription factor 31 | ||||
| STRING | MLOC_34885.1 | 1e-167 | (Hordeum vulgare) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP229 | 38 | 296 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G47600.1 | 1e-102 | myb domain protein 94 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



