 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | HL.SW.v1.0.G044380.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Cannabaceae; Humulus
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 367aa MW: 42588.6 Da PI: 8.8452 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 20.1 | 1.7e-06 | 64 | 85 | 2 | 23 |
EETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 2 kCpdCgksFsrksnLkrHirtH 23
+C+ Cg +F+ +Lk+H+++H
HL.SW.v1.0.G044380.1 64 TCEECGLTFKKAAYLKQHMQSH 85
6*******************99 PP
| |||||||
| 2 | zf-C2H2 | 16.2 | 2.9e-05 | 91 | 115 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
++C+ dC s++rk++L+rH+ H
HL.SW.v1.0.G044380.1 91 HVCSvdDCRSSYRRKDHLTRHLLIH 115
799*******************998 PP
| |||||||
| 3 | zf-C2H2 | 12.5 | 0.00044 | 120 | 145 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
+kCp +C+ Fs + n krH r H
HL.SW.v1.0.G044380.1 120 FKCPieNCKIEFSIQANVKRHVREkH 145
89*********************988 PP
| |||||||
| 4 | zf-C2H2 | 16.9 | 1.8e-05 | 160 | 184 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
++C+ Cgk F +s L++H ++H
HL.SW.v1.0.G044380.1 160 HVCQevGCGKAFAYPSRLRKHEQSH 184
789999***************9998 PP
| |||||||
| 5 | zf-C2H2 | 14.7 | 8.7e-05 | 252 | 275 | 3 | 23 |
ET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 3 Cp..dCgksFsrksnLkrHirt.H 23
C C +Fs+ksnL++H++ H
HL.SW.v1.0.G044380.1 252 CLykGCLHTFSTKSNLNQHLKAvH 275
66679***************9877 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00355 | 27 | 17 | 39 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 13.297 | 63 | 90 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0055 | 63 | 85 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 8.5E-6 | 64 | 84 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 2.19E-10 | 64 | 115 | No hit | No description |
| PROSITE pattern | PS00028 | 0 | 65 | 85 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 4.2E-12 | 85 | 117 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 9.494 | 91 | 120 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.037 | 91 | 115 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 93 | 115 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 2.64E-7 | 101 | 155 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 3.0E-4 | 118 | 143 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.448 | 120 | 150 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.019 | 120 | 145 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 122 | 145 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 1.7E-8 | 154 | 184 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 1.15E-5 | 156 | 186 | No hit | No description |
| PROSITE profile | PS50157 | 11.136 | 160 | 189 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0034 | 160 | 184 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 162 | 184 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 6.9 | 192 | 217 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 194 | 217 | IPR007087 | Zinc finger, C2H2 |
| CDD | cd02980 | 0.0089 | 212 | 257 | No hit | No description |
| SMART | SM00355 | 12 | 220 | 241 | IPR015880 | Zinc finger, C2H2-like |
| SuperFamily | SSF57667 | 7.95E-8 | 231 | 292 | No hit | No description |
| SMART | SM00355 | 0.0088 | 250 | 275 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.697 | 250 | 280 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 1.3E-4 | 251 | 277 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 252 | 275 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 9.8E-6 | 278 | 302 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 6.8 | 281 | 307 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 283 | 307 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008097 | Molecular Function | 5S rRNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0080084 | Molecular Function | 5S rDNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 367 aa Download sequence Send to blast |
MVEAGEERPI FRDIRRYFCD YCGICRSKKT LISSHILTHH KEELDKVKAE GESSTEGVEE 60 SKNTCEECGL TFKKAAYLKQ HMQSHSLERP HVCSVDDCRS SYRRKDHLTR HLLIHKGKLF 120 KCPIENCKIE FSIQANVKRH VREKHNEDRP STSTEREKQH VCQEVGCGKA FAYPSRLRKH 180 EQSHVKLESI EALCVEPGCM KIFTNEQCLR DHIQLCHQHI TCEVCGSKHL KKNMKRHLRS 240 HEEKVSVESI KCLYKGCLHT FSTKSNLNQH LKAVHFNKKP YVCGFSGCGK RFAYKHVRDN 300 HEKRSCHIYA HGDFEEADEQ FRSRPRGGRK RKYPSFVDLL VRKRITPRND LDGSFECLDS 360 FSCGGE* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2i13_A | 4e-14 | 60 | 244 | 19 | 187 | Aart |
| 2i13_B | 4e-14 | 60 | 244 | 19 | 187 | Aart |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 229 | 235 | LKKNMKR |
| 2 | 323 | 331 | RPRGGRKRK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Essential protein (PubMed:22353599). Isoform 1 is a transcription activator the binds both 5S rDNA and 5S rRNA and stimulates the transcription of 5S rRNA gene (PubMed:12711688, PubMed:22353599). Isoform 1 regulates 5S rRNA levels during development (PubMed:22353599). {ECO:0000269|PubMed:12711688, ECO:0000269|PubMed:22353599}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00229 | DAP | Transfer from AT1G72050 | Download |
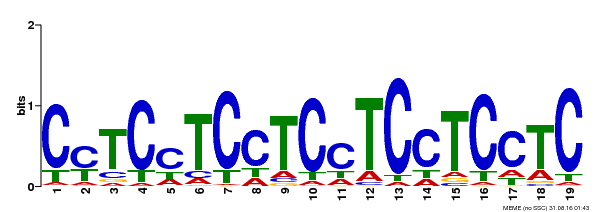
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015867533.1 | 0.0 | transcription factor IIIA isoform X1 | ||||
| Refseq | XP_024924686.1 | 0.0 | transcription factor IIIA isoform X1 | ||||
| Swissprot | Q84MZ4 | 1e-139 | TF3A_ARATH; Transcription factor IIIA | ||||
| TrEMBL | A0A2P5CW95 | 0.0 | A0A2P5CW95_PARAD; TFIIH C1-like domain containing protein | ||||
| STRING | XP_010104482.1 | 0.0 | (Morus notabilis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF3069 | 33 | 70 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G72050.2 | 1e-141 | transcription factor IIIA | ||||



