 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | HL.SW.v1.0.G044101.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Cannabaceae; Humulus
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 842aa MW: 91293.1 Da PI: 5.9818 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 66.1 | 4.7e-21 | 130 | 185 | 1 | 56 |
TT--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 1 rrkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ +++t++q++eLe+lF+++++p++++r eL+k+l L++rqVk+WFqNrR+++k
HL.SW.v1.0.G044101.1 130 KKRYHRHTPQQIQELEALFKECPHPDEKQRLELSKRLCLETRQVKFWFQNRRTQMK 185
688999***********************************************999 PP
| |||||||
| 2 | START | 209.1 | 1.7e-65 | 335 | 560 | 1 | 206 |
HHHHHHHHHHHHHHHHC-TT-EEEE....EXCCTTEEEEEEESSS......SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-S... CS
START 1 elaeeaaqelvkkalaeepgWvkss....esengdevlqkfeeskv.....dsgealrasgvvdmvlallveellddkeqWdetla... 77
ela++a++elvk+a+ eep+W ks+ + +n++e++++f + + + ++a+r+sg+v+ ++ lve+l++++ +W e+++
HL.SW.v1.0.G044101.1 335 ELALAAMDELVKMAQTEEPLWIKSYeggrDVLNQEEYMRSFNPCIGmkpsgFVTDASRESGIVIINSLALVETLMESS-RWAEMFPcvi 422
5899*************************************99888999999**************************.********** PP
.EEEEEEEECTT......EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--..-TTSEE-EESSEEEEEEEE CS
START 78 .kaetlevissg......galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe.sssvvRaellpSgiliepk 157
+ +t++vissg galqlm aelq+lsplvp R++ f+R+++q+ +g+w++vdvS+d +++++ +s++ +++lpSg++++++
HL.SW.v1.0.G044101.1 423 aRTSTTDVISSGmggtrnGALQLMHAELQVLSPLVPvREVNFLRFCKQHAEGVWAVVDVSIDTIRDNSSgPPSFINCRRLPSGCVVQDM 511
********************************************************************999****************** PP
CTCEEEEEEEE-EE--SSXXHHHHHHHHHHHHHHHHHHHHHHTXXXXXX CS
START 158 snghskvtwvehvdlkgrlphwllrslvksglaegaktwvatlqrqcek 206
+ g+skvtwveh++++++++h+l+r+l++sg+ +ga++wvatlqrqce+
HL.SW.v1.0.G044101.1 512 PSGYSKVTWVEHAEYDESQVHQLYRPLLSSGMGFGAQRWVATLQRQCEC 560
***********************************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.03E-20 | 112 | 187 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 7.5E-22 | 117 | 187 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.216 | 127 | 187 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 9.9E-18 | 128 | 191 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.38E-18 | 129 | 187 | No hit | No description |
| Pfam | PF00046 | 1.4E-18 | 130 | 185 | IPR001356 | Homeobox domain |
| PROSITE pattern | PS00027 | 0 | 162 | 185 | IPR017970 | Homeobox, conserved site |
| PROSITE profile | PS50848 | 43.618 | 326 | 563 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 1.65E-34 | 328 | 560 | No hit | No description |
| CDD | cd08875 | 8.70E-122 | 330 | 559 | No hit | No description |
| Pfam | PF01852 | 3.4E-58 | 335 | 560 | IPR002913 | START domain |
| SMART | SM00234 | 1.5E-50 | 335 | 560 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 9.89E-24 | 589 | 759 | No hit | No description |
| SuperFamily | SSF55961 | 9.89E-24 | 794 | 834 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009827 | Biological Process | plant-type cell wall modification | ||||
| GO:0042335 | Biological Process | cuticle development | ||||
| GO:0043481 | Biological Process | anthocyanin accumulation in tissues in response to UV light | ||||
| GO:0048765 | Biological Process | root hair cell differentiation | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 842 aa Download sequence Send to blast |
MSFGGFLDNS TSTSGGGARL VAEIPYSSTS NDNIGNGNDN NHIMPSTAIA QPRLITQSLT 60 KSMFNSPGLS LALQTNRDGQ GDMIRNMAES FEPGSGRRSR DEEHESRSGS DNMDGGSGDD 120 QDATDKPPRK KRYHRHTPQQ IQELEALFKE CPHPDEKQRL ELSKRLCLET RQVKFWFQNR 180 RTQMKTQLER HENSLLRQEN DKLRAENMSI RDAMRNPMCT NCGGPAIIGE ISLEEQHLRI 240 ENARLKDELE RVCSLAGKFL GRPISALATS MAPPLPSSAL ELGVGSNGFG GLTSVTTTMP 300 DFGISSNTLA VLPQSRSQGG VQVLDRSIER SMYLELALAA MDELVKMAQT EEPLWIKSYE 360 GGRDVLNQEE YMRSFNPCIG MKPSGFVTDA SRESGIVIIN SLALVETLME SSRWAEMFPC 420 VIARTSTTDV ISSGMGGTRN GALQLMHAEL QVLSPLVPVR EVNFLRFCKQ HAEGVWAVVD 480 VSIDTIRDNS SGPPSFINCR RLPSGCVVQD MPSGYSKVTW VEHAEYDESQ VHQLYRPLLS 540 SGMGFGAQRW VATLQRQCEC LAILMSSTVP TRDHTAGITP GGRRSMLKLA QRMTDNFCAG 600 VCASTVHKWN KLNAGNVDED VRVMTRKSVD DPGEPPGIVL SAATSVWLPV SPQRLFDFLR 660 DESLRSEWDI LSNGGPMQEM AHIAKGQNHG NCVSLLRASA MNTNQSSMLI LQETCIDEAG 720 SLVVYAPVDI PAMHVVMNGG DSAYVALLPS GFAIVPDGPG SRGPNMGPTA NGGGNNNGGS 780 GSNGNGGDGS HRVGGSLLTV AFQILVNSLP TAKLTVESVE TVNNLISCTV QKIKAALQCE 840 S* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of the tissue-specific accumulation of anthocyanins and in cellular organization of the primary root. {ECO:0000269|PubMed:10402424}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00421 | DAP | Transfer from AT4G00730 | Download |
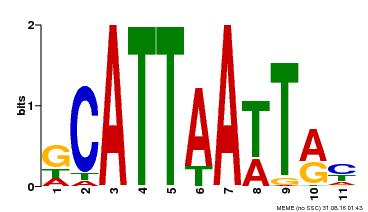
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024018220.1 | 0.0 | homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| Swissprot | Q0WV12 | 0.0 | ANL2_ARATH; Homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| TrEMBL | A0A2P5CNQ4 | 0.0 | A0A2P5CNQ4_TREOI; Octamer-binding transcription factor | ||||
| STRING | XP_010091553.1 | 0.0 | (Morus notabilis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1192 | 34 | 106 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G00730.1 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



