 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | HL.SW.v1.0.G038666.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Cannabaceae; Humulus
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 206aa MW: 22727.2 Da PI: 5.0912 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 58.7 | 1.5e-18 | 19 | 68 | 2 | 55 |
AP2 2 gykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
+ykGVr +k +g+Wv+eIr p++ r r++lg+++tae+Aa+a +aa +l+g
HL.SW.v1.0.G038666.1 19 RYKGVRKRK-WGKWVSEIRLPNS---RERIWLGSYDTAEKAARAFDAALFCLRG 68
69****999.**********933...5*************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF00847 | 2.1E-11 | 19 | 68 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 22.563 | 19 | 76 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 6.18E-30 | 19 | 78 | No hit | No description |
| SMART | SM00380 | 1.2E-34 | 19 | 82 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 8.5E-21 | 19 | 77 | IPR016177 | DNA-binding domain |
| Gene3D | G3DSA:3.30.730.10 | 7.0E-30 | 19 | 78 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 4.3E-10 | 20 | 31 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 4.3E-10 | 42 | 58 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 206 aa Download sequence Send to blast |
MVKQSPIEKN PESERSDLRY KGVRKRKWGK WVSEIRLPNS RERIWLGSYD TAEKAARAFD 60 AALFCLRGRT AKFNFPNSPP EIAGGRSLTP TEIQAAAARF ANSEAANGRH SGRTVPEIIQ 120 MESPSPSVSE GSGMAHLDSD VATNLSCMDL FSNLGTGNYA TEYGLFPGFD DLTGDFFSPP 180 LPTIDYGEDN SFDGYLHQDS FLWNF* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5wx9_A | 8e-16 | 19 | 77 | 14 | 73 | Ethylene-responsive transcription factor ERF096 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 20 | 26 | KGVRKRK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probably acts as a transcriptional activator. Binds to the GCC-box pathogenesis-related promoter element. May be involved in the regulation of gene expression by stress factors and by components of stress signal transduction pathways (By similarity). {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00151 | DAP | Transfer from AT1G19210 | Download |
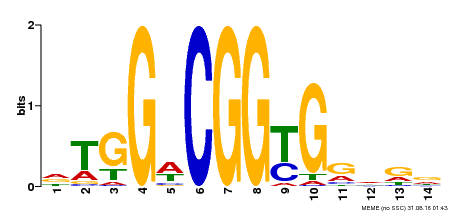
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KF678386 | 3e-45 | KF678386.1 Morus notabilis dehydration responsive element binding transcription factor (DREB5H) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010099089.1 | 5e-99 | ethylene-responsive transcription factor ERF017 | ||||
| Swissprot | Q84QC2 | 1e-60 | ERF17_ARATH; Ethylene-responsive transcription factor ERF017 | ||||
| TrEMBL | A0A2P5EGG1 | 1e-126 | A0A2P5EGG1_TREOI; AP2/ERF domain containing protein | ||||
| STRING | XP_010099089.1 | 2e-98 | (Morus notabilis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1479 | 33 | 99 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G19210.1 | 3e-55 | ERF family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



