 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | HL.SW.v1.0.G029945.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Cannabaceae; Humulus
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 424aa MW: 47600.2 Da PI: 8.2501 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 101.1 | 7.5e-32 | 75 | 128 | 2 | 55 |
G2-like 2 prlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
prlrWtp+LH rFv+ave+LGG+e+AtPk +l+lm++kgL+++hvkSHLQ+YR+
HL.SW.v1.0.G029945.1 75 PRLRWTPDLHLRFVHAVERLGGQERATPKLVLQLMSIKGLSIAHVKSHLQMYRS 128
8****************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 1.0E-29 | 71 | 129 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 12.282 | 71 | 131 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.97E-14 | 73 | 129 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.4E-22 | 75 | 130 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.0E-8 | 76 | 127 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 424 aa Download sequence Send to blast |
MINMVVESRT SDSECSKTSF EYEDESGEEN EDESTKPNNK DTNNGGLSSS NSTVEESDKV 60 KSSSSVRPYV RSKMPRLRWT PDLHLRFVHA VERLGGQERA TPKLVLQLMS IKGLSIAHVK 120 SHLQMYRSKK IDDAGQVIGS HDHHHRHLLD CGDRNIFNLS QLPMLQGYKQ TSHSSNFRYG 180 FDDPSWTSFG NLRHSRAGFY ALSAEAERKI LASNWGSSTN ACNFPSKNNI SSTISSNLIN 240 HDQHSSWRKA HNKLTIKDND HDHHNNHLQH RPSSTPHQTQ FNTPLIDLNS IAHLQVNKAK 300 ELTNFLSSNN NVTHSDTVLN TTTTTTSTST SMKRKASSDS NNHHHHHQLD LDLSLRLTTT 360 SSDVVMDIDR DHKRKRSSGD EVDSSLLSLS LNSSNSSSKR SKNDDHHHGE MQARRASTLD 420 LTI* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 2e-17 | 76 | 130 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 2e-17 | 76 | 130 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 2e-17 | 76 | 130 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 2e-17 | 76 | 130 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 2e-17 | 76 | 130 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 2e-17 | 76 | 130 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 2e-17 | 76 | 130 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 2e-17 | 76 | 130 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00297 | DAP | Transfer from AT2G38300 | Download |
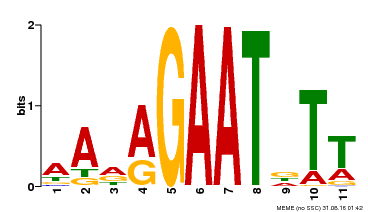
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010092813.1 | 1e-110 | two-component response regulator ORR22 | ||||
| TrEMBL | W9R2E6 | 1e-109 | W9R2E6_9ROSA; Putative Myb family transcription factor | ||||
| STRING | XP_010092813.1 | 1e-110 | (Morus notabilis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF2378 | 34 | 82 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G38300.1 | 2e-52 | G2-like family protein | ||||



