 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | HL.SW.v1.0.G023714.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Cannabaceae; Humulus
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1494aa MW: 167172 Da PI: 8.831 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 12.9 | 0.00033 | 1403 | 1425 | 3 | 23 |
ET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 3 Cp..dCgksFsrksnLkrHirtH 23
Cp Cgk F ++ +L++H r+H
HL.SW.v1.0.G023714.1 1403 CPvkGCGKKFFSHKYLVQHRRVH 1425
9999*****************99 PP
| |||||||
| 2 | zf-C2H2 | 11.8 | 0.00073 | 1461 | 1487 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH..T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt..H 23
y+C Cg++F+ s++ rH r+ H
HL.SW.v1.0.G023714.1 1461 YVCAepGCGQTFRFVSDFSRHKRKtgH 1487
899999****************99666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00545 | 1.1E-17 | 23 | 64 | IPR003349 | JmjN domain |
| PROSITE profile | PS51183 | 14.853 | 24 | 65 | IPR003349 | JmjN domain |
| Pfam | PF02375 | 3.1E-14 | 25 | 58 | IPR003349 | JmjN domain |
| SMART | SM00558 | 3.1E-45 | 196 | 370 | IPR003347 | JmjC domain |
| PROSITE profile | PS51184 | 31.69 | 199 | 370 | IPR003347 | JmjC domain |
| SuperFamily | SSF51197 | 9.89E-23 | 210 | 372 | No hit | No description |
| Pfam | PF02373 | 1.6E-33 | 229 | 352 | IPR003347 | JmjC domain |
| SMART | SM00355 | 28 | 1378 | 1400 | IPR015880 | Zinc finger, C2H2-like |
| SuperFamily | SSF57667 | 6.07E-6 | 1401 | 1437 | No hit | No description |
| PROSITE profile | PS50157 | 12.445 | 1401 | 1430 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0045 | 1401 | 1425 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 1.2E-5 | 1403 | 1429 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 1403 | 1425 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 1.3E-7 | 1430 | 1455 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 10.637 | 1431 | 1460 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0017 | 1431 | 1455 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1433 | 1455 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 2.92E-9 | 1441 | 1485 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 2.6E-9 | 1456 | 1484 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.032 | 1461 | 1492 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.62 | 1461 | 1487 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1463 | 1487 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0040010 | Biological Process | positive regulation of growth rate | ||||
| GO:0045815 | Biological Process | positive regulation of gene expression, epigenetic | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0071558 | Molecular Function | histone demethylase activity (H3-K27 specific) | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1494 aa Download sequence Send to blast |
MAASALGSDP SPEVFSWLKT LPLAPEYHPT LTEFQDPISY IFKIEKEASK YGICKIVPPV 60 SPSPKKTVIA NLNKSLAARA GYSEGSNPKT PPTFTTRQQQ IGFCPRKPRP VQRPVWQSGE 120 QYTFQEFETK AKGFEKGFLK KCGKKGGGLS PLEIETLYWK ATVDKPFSVE YANDMPGSAF 180 VPVSAKKSRE AGESVTLGET AWNMRAVSRS KGSLLRFMKE EIPGVTSPMV YVAMLFSWFA 240 WHVEDHDLHS LNYLHMGASK TWYGVPKEAA VAFEEVIRVH GYGAEINPLV TFSILGEKTT 300 VMSPEVFVKE GVPCCRLVQN AGEFVVTFPR AYHTGFSHGE RPRRFNCAEA ANIATPEWLR 360 VANDAAIRRA SINYPPMVSH FQLLYDLALA LCSRIPVSIS SEPRSSRLKD KKKGEGETVV 420 KELFVQDVLK NNDILRVLGK GSPIVLLPRT SSDISVCSKL RVGSHLRLNT NVSLSPSISR 480 EEIKTSRSLI SDDLTIDRKQ EVDQEKVFYS VKGKLASLCE GSWVPSRGNK RTSTSNFKTP 540 IKNVEEKSNV DSDRLSEQRL FSCVTCGILS FACVAIVQPR DPAARYLMTA DCSFFNDWVV 600 NAGVASNVFP ISSVDQTASK QNTYTAWADN CAPLDSYEGP VPSISNQVQT EDQNNEVLSN 660 SEAHEATSAL GLLALHYGNS SDSEEDQARE DDSVYDKETN TTNCSPESKY RCESSSLSFQ 720 NSLSDAAGDD GRSPREIDLG EERASQNADF HKENGHNRDN VKYSSRQNFD CSVDASNNVA 780 STETNCLVGK IRDPMIVSQA CSADTYDAEA TRFCKATVPM KNDVVPFVPL SDEESSRMHV 840 FCLEHAVEVE QQLRQIGGVH ILLLCHPDYP KIETEAKSIA EELGINCPLN DTTFRDATKD 900 DQEKIQSALD SEEAMPKNGD WAVKLGINLF YSANLSRSPL YSKQMPYNSV IYNAFGRGSP 960 TSSSMKSDGF GRRSVKKKVV AGKWCGKVWM SNQVHPFLVK RNPEEEEERG ILAWAVPDEK 1020 PDRNHDSPHR NSNTMVTKKY VRKRKMKVEN ESIKKAKCIK GEDIVSENSM DGDSHEHHRR 1080 SLGNNQSTHT GVGSAKKGKH IQMEDAVSDD SMYVSLKQCK RTLRSKVTTS VETDDIVSDD 1140 SLGVISNYRN KKNPRREQAK QVKNLGIQDA VSDGSLDSES NQRRRRIPKS KQAKFMEEED 1200 SASDDFPGTN LYKQQTRIYK SKYAKSIERE DEVSDEPEEA YACHEDKRVL RSKQTKSTMQ 1260 PKMKQGTGRH VRQPSSQPVK QGTRQLAKKE TPKLKQQTPR LRNNQCDQNT SHICDDEEEE 1320 GGPSTRLRKR TPKPQKTAGP KKKEQQQQTS NKKAKNASAV KAQAGRGDAK FKDEEAEYVC 1380 DVEGCTMSFG VKHELVLHKR NICPVKGCGK KFFSHKYLVQ HRRVHMDDRP LRCPWKGCKM 1440 TFKWAWARTE HIRVHTGARP YVCAEPGCGQ TFRFVSDFSR HKRKTGHSTK KGR* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6ip0_A | 6e-74 | 17 | 391 | 8 | 353 | Transcription factor jumonji (Jmj) family protein |
| 6ip4_A | 6e-74 | 17 | 391 | 8 | 353 | Arabidopsis JMJ13 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 971 | 977 | RRSVKKK |
| 2 | 1038 | 1044 | KYVRKRK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Histone demethylase that demethylates 'Lys-27' (H3K27me) of histone H3. Demethylates both tri- (H3K27me3) and di-methylated (H3K27me2) H3K27me (PubMed:21642989, PubMed:27111035). Demethylates also H3K4me3/2 and H3K36me3/2 in an in vitro assay (PubMed:20711170). Involved in the transcriptional regulation of hundreds of genes regulating developmental patterning and responses to various stimuli (PubMed:18467490). Binds DNA via its four zinc fingers in a sequence-specific manner, 5'-CTCTGYTY-3', to promote the demethylation of H3K27me3 and the regulation of organ boundary formation (PubMed:27111034, PubMed:27111035). Involved in the regulation of flowering time by repressing FLOWERING LOCUS C (FLC) expression (PubMed:15377760). Interacts with the NF-Y complexe to regulate SOC1 (PubMed:25105952). Mediates the recruitment of BRM to its target loci (PubMed:27111034). {ECO:0000269|PubMed:15377760, ECO:0000269|PubMed:18467490, ECO:0000269|PubMed:20711170, ECO:0000269|PubMed:21642989, ECO:0000269|PubMed:25105952, ECO:0000269|PubMed:27111034, ECO:0000269|PubMed:27111035}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
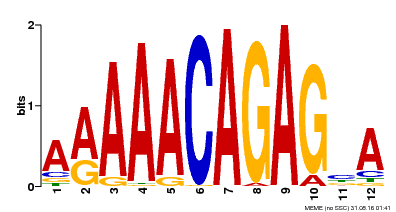
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010101942.1 | 0.0 | lysine-specific demethylase REF6 | ||||
| Swissprot | Q9STM3 | 0.0 | REF6_ARATH; Lysine-specific demethylase REF6 | ||||
| TrEMBL | A0A2P5EIB2 | 0.0 | A0A2P5EIB2_TREOI; TFIIH C1-like domain containing protein | ||||
| STRING | XP_010101942.1 | 0.0 | (Morus notabilis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF5648 | 32 | 50 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 0.0 | relative of early flowering 6 | ||||



