 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | HL.SW.v1.0.G023154.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Cannabaceae; Humulus
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 623aa MW: 69316.8 Da PI: 6.6021 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 40.4 | 5.2e-13 | 453 | 499 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCC.C...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPk.askapskKlsKaeiLekAveYIksLq 55
+h e+Er+RR+++N++f Lr ++P+ + K++Ka+ L A+ YI++Lq
HL.SW.v1.0.G023154.1 453 NHVEAERQRREKLNQRFYALRAVVPNiS------KMDKASLLGDAIAYINELQ 499
799***********************66......******************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 2.1E-56 | 53 | 241 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| PROSITE profile | PS50888 | 17.047 | 449 | 498 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SuperFamily | SSF47459 | 2.49E-18 | 452 | 516 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 1.38E-14 | 452 | 503 | No hit | No description |
| Gene3D | G3DSA:4.10.280.10 | 1.4E-18 | 453 | 512 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 1.8E-10 | 453 | 499 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 1.0E-16 | 455 | 504 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 623 aa Download sequence Send to blast |
MKTELGFGGS GVWNDEDKTM VQAVLGTRAF DYLITSSVTN ENLLMAVGSD ENLQNKLSDL 60 VERPNASNFS WNYAIFWQIS RSKSGDWVLG WGDGSCREPR EGEESEATRF FNLRLEDETQ 120 QRMRKRVLQK LHNLFGGSDE DNYALGLDRV TDTEIFFLAS MYFSFPRGEG GPGKCYASGK 180 HLWLLDALKS SSDYCVRSFL AKSAGIQTIV LVPTEVGVVE LGSVRSVGES LELLQSVRSL 240 FSTQSSLVRV SKPMAMLPMM AEKRDENVNL TNSGILERNE SVGVHKIFGQ DLNAGAVARP 300 HYREKLAIRK LEERPWDLYP NGNGSRIAFS SPRNGIPGST WPHSLQGVKQ QGNPTEIYAS 360 QSPGSNNIQE IVNGVREDFR LNNYQPQKQV QMQIDFSGAT SRSTVINHRP TSAESEHSDV 420 EASCKEERPS TADERRPRKR GRKPANGREE PLNHVEAERQ RREKLNQRFY ALRAVVPNIS 480 KMDKASLLGD AIAYINELQA KLKAMEAERE KVGSSSTAAA METNNPDMEI QSRVPDVDVK 540 VVNDEVIVRV SCHLDSHPAS RVIQAFKEAQ ITVVDSKFTT VNDTVFHDFV IKSQGSEQLT 600 KEKLVAAFSI EANSLRTLSS VG* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5gnj_A | 7e-30 | 447 | 522 | 2 | 77 | Transcription factor MYC2 |
| 5gnj_B | 7e-30 | 447 | 522 | 2 | 77 | Transcription factor MYC2 |
| 5gnj_E | 7e-30 | 447 | 522 | 2 | 77 | Transcription factor MYC2 |
| 5gnj_F | 7e-30 | 447 | 522 | 2 | 77 | Transcription factor MYC2 |
| 5gnj_G | 7e-30 | 447 | 522 | 2 | 77 | Transcription factor MYC2 |
| 5gnj_I | 7e-30 | 447 | 522 | 2 | 77 | Transcription factor MYC2 |
| 5gnj_M | 7e-30 | 447 | 522 | 2 | 77 | Transcription factor MYC2 |
| 5gnj_N | 7e-30 | 447 | 522 | 2 | 77 | Transcription factor MYC2 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 434 | 442 | RRPRKRGRK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that negatively regulates jasmonate (JA) signaling (PubMed:30610166). Negatively regulates JA-dependent response to wounding, JA-induced expression of defense genes, JA-dependent responses against herbivorous insects, and JA-dependent resistance against Botrytis cinerea infection (PubMed:30610166). Plays a positive role in resistance against the bacterial pathogen Pseudomonas syringae pv tomato DC3000 (PubMed:30610166). {ECO:0000269|PubMed:30610166}. | |||||
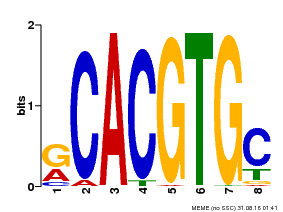
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00100 | PBM | Transfer from AT1G01260 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by wounding, feeding with herbivorous insects, infection with the fungal pathogen Botrytis cinerea and infection with the bacterial pathogen Pseudomonas syringae pv tomato DC3000. {ECO:0000269|PubMed:30610166}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024024130.1 | 0.0 | LOW QUALITY PROTEIN: transcription factor bHLH13 | ||||
| Swissprot | A0A3Q7ELQ2 | 0.0 | MTB1_SOLLC; Transcription factor MTB1 | ||||
| TrEMBL | A0A2P5FHN3 | 0.0 | A0A2P5FHN3_TREOI; Basic helix-loop-helix transcription factor | ||||
| STRING | XP_010100724.1 | 0.0 | (Morus notabilis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF5448 | 32 | 54 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G01260.3 | 0.0 | bHLH family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



