 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | HL.SW.v1.0.G016605.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Cannabaceae; Humulus
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 881aa MW: 101245 Da PI: 7.298 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 84.4 | 1.9e-26 | 89 | 191 | 1 | 91 |
FAR1 1 kfYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkk.........tekerr.......traetrtgCkaklkvkkekd 73
+fY+eYA+++GF++ +++s++sk+++e+++++f Cs++g+++e +k+ + + +r+ ++t+Cka+++vk++ d
HL.SW.v1.0.G016605.1 89 SFYQEYARSMGFNTAIQNSRRSKTSREFIDAKFACSRYGTKREYDKSfnrprsrqsK----QdpdhltgRRSCSKTDCKASMHVKRRPD 173
5*******************************************9999998865440....24556666999***************** PP
FAR1 74 gkwevtkleleHnHelap 91
gkw+++++++eHnHel p
HL.SW.v1.0.G016605.1 174 GKWVIHNFVKEHNHELLP 191
***************975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 3.0E-24 | 89 | 191 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 4.5E-31 | 290 | 381 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 9.446 | 569 | 605 | IPR007527 | Zinc finger, SWIM-type |
| Pfam | PF04434 | 1.1E-5 | 578 | 604 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 1.1E-8 | 580 | 607 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009585 | Biological Process | red, far-red light phototransduction | ||||
| GO:0010218 | Biological Process | response to far red light | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 881 aa Download sequence Send to blast |
MDIDLRLPSG EHDKEGEEST AIDNMLDGEE KLHNGDIISG NMVDIGDEVQ TEDGGDLNSP 60 TANMVVFKED TNLEPLSGME FESHGEAYSF YQEYARSMGF NTAIQNSRRS KTSREFIDAK 120 FACSRYGTKR EYDKSFNRPR SRQSKQDPDH LTGRRSCSKT DCKASMHVKR RPDGKWVIHN 180 FVKEHNHELL PAQAVSEQTR KMYAAMARQF AEYKNVVGLR NDPKNPFEKG RNLAVEAGDL 240 KILLDFFMQM QNVNANFFYA VDVGEDQRIK NLFWVDAKSR HDYANFSDAV SFDTTYIRNK 300 YKMPLALFVG VNQHYQFMLL GCALVSDESV TTFSWLMQTW LKAMGGQVPK VIITDHDKVL 360 KSVIFEVFPT AHHCFCLWHI LVRVSENLGH VIKKHENFMA KFEKCIYRSW TVEEFEKRWW 420 KILDKCELRG DEWIHSLYED RKQWVPSFMR DALLAGMSTV QRSDSVNYYF DKYVHKKTTV 480 QEFLRQHEAI LQDRYEEEAK ADSDTWNKQP TLKSPSPLEK SVSGVYTHAV FKKFQVEVLG 540 AVACIPKKER QDEMSIIFRV QDFEKEQDFQ VSWNEMKSEV SCLCRLFEYK GYLCRHALIV 600 LQMCGLSVIP PQYILKRWTR DAKNRHVTGE ESGQLQSRVQ RYNDLCQRAM KLSEEGSLSQ 660 ESYSIACRAL DETFGNCTSV NNSSRSLVDA GTSATHGLLC MEEDSQSKNI GKTNKKKNPT 720 KKRKVSFEPD VMTVGTQDSL QQMDKLNSRA VTLDSYYGAQ QNVQGMVQLN LMAPTRDNYY 780 GNQQTIQGLG QLNSIAPSHD GYYSAQQSMH GLGQMDFFRG TAPGFTYGIR DDPNVRAASL 840 HDDASRHGYR RHKSHSFEEK IGSCLVWKDG SRENLPVWHL * |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
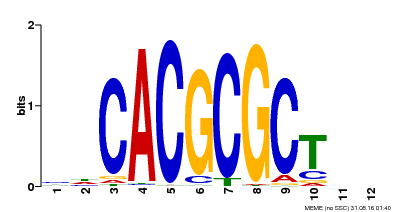
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. When associated with PHYA, protects it from being recognized and degraded by the COP1/SPA complex. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:17012604, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:18715961, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00078 | ChIP-seq | Transfer from AT3G22170 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. Up-regulated by white light. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024019194.1 | 0.0 | protein FAR-RED ELONGATED HYPOCOTYL 3 isoform X1 | ||||
| Swissprot | Q9LIE5 | 0.0 | FHY3_ARATH; Protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| TrEMBL | A0A251Q297 | 0.0 | A0A251Q297_PRUPE; Uncharacterized protein | ||||
| STRING | EMJ16129 | 0.0 | (Prunus persica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF40 | 30 | 530 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G22170.2 | 0.0 | far-red elongated hypocotyls 3 | ||||



