 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | HL.SW.v1.0.G015938.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Cannabaceae; Humulus
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 332aa MW: 37403.2 Da PI: 4.8425 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 59.4 | 5.8e-19 | 57 | 110 | 3 | 56 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k+++++ +q+++Le+ Fe ++++ e++ +LA++lgL+ rqV vWFqNrRa++k
HL.SW.v1.0.G015938.1 57 KKRRLSVDQVKALEKNFEVENKLEPERKVKLAQELGLQPRQVAVWFQNRRARWK 110
566899***********************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 130.1 | 9.1e-42 | 56 | 148 | 1 | 93 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLre 89
ekkrrls +qvk+LE++Fe e+kLeperKv+la+eLglqprqvavWFqnrRAR+ktkqlE+dy +Lk++ydalk + ++L++++e+L +
HL.SW.v1.0.G015938.1 56 EKKRRLSVDQVKALEKNFEVENKLEPERKVKLAQELGLQPRQVAVWFQNRRARWKTKQLERDYGHLKANYDALKLSYDTLKHDNETLVK 144
69**************************************************************************************9 PP
HD-ZIP_I/II 90 elke 93
e++e
HL.SW.v1.0.G015938.1 145 EIRE 148
9986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 2.4E-19 | 44 | 114 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.443 | 52 | 112 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 6.3E-18 | 55 | 116 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.50E-16 | 57 | 113 | No hit | No description |
| Pfam | PF00046 | 3.7E-16 | 57 | 110 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 8.4E-20 | 59 | 119 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 9.6E-6 | 83 | 92 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 87 | 110 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 9.6E-6 | 92 | 108 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 2.9E-17 | 112 | 154 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009637 | Biological Process | response to blue light | ||||
| GO:0030308 | Biological Process | negative regulation of cell growth | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0048510 | Biological Process | regulation of timing of transition from vegetative to reproductive phase | ||||
| GO:0048573 | Biological Process | photoperiodism, flowering | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 332 aa Download sequence Send to blast |
MKRLSSDDSL GALMAVCPTT EEHSPSSNHV YGREFRSMLD GLDEEGCVEE SGHVSEKKRR 60 LSVDQVKALE KNFEVENKLE PERKVKLAQE LGLQPRQVAV WFQNRRARWK TKQLERDYGH 120 LKANYDALKL SYDTLKHDNE TLVKEIRELK SKLQDEKSES NNLSVKEELH IGPTESENKH 180 AAHPDHHILP SEECKELNFE NLNDTNNGVV LEVSMFPDFN KDGSSDSDSS AILNDDINLN 240 TNISSSPNGA ISSTGVLQNH QLMSSPSPVA SASLNFNCSS SSSNCFQFQK AASFQPQFVK 300 IEEHNFFAEE ACNFFSDDQA PTLQWYCSDQ W* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 104 | 112 | RRARWKTKQ |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00051 | PBM | Transfer from AT4G40060 | Download |
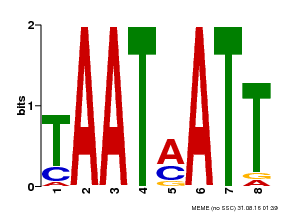
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_016651837.1 | 1e-155 | PREDICTED: homeobox-leucine zipper protein ATHB-6-like isoform X2 | ||||
| TrEMBL | A0A2P5AY58 | 1e-172 | A0A2P5AY58_PARAD; Homeobox protein | ||||
| STRING | XP_008235150.1 | 1e-152 | (Prunus mume) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1868 | 34 | 89 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G40060.1 | 2e-69 | homeobox protein 16 | ||||



