 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | HL.SW.v1.0.G007058.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Cannabaceae; Humulus
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 763aa MW: 82622.5 Da PI: 7.1901 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 86.9 | 2e-27 | 193 | 247 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
k +++WtpeLH+rFv+aveqL G +kA+P++ilelm++++Lt+++++SHLQkYR++
HL.SW.v1.0.G007058.1 193 KVKVDWTPELHRRFVQAVEQL-GVDKAVPSRILELMGIDCLTRHNIASHLQKYRSH 247
5689*****************.********************************86 PP
| |||||||
| 2 | G2-like | 85.2 | 6.6e-27 | 475 | 526 | 4 | 56 |
G2-like 4 lrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
++WtpeLH+rFv+aveqL G +kA+P++ilelm++++Lt+++++SHLQkYR++
HL.SW.v1.0.G007058.1 475 VDWTPELHRRFVQAVEQL-GVDKAVPSRILELMGIDCLTRHNIASHLQKYRSH 526
79****************.********************************86 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 18.152 | 190 | 249 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.6E-26 | 191 | 250 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 8.17E-19 | 191 | 250 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 4.0E-26 | 193 | 247 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 9.7E-8 | 196 | 245 | IPR001005 | SANT/Myb domain |
| PROSITE profile | PS51294 | 14.687 | 469 | 528 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.52E-18 | 471 | 529 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 1.7E-25 | 472 | 529 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.0E-25 | 473 | 526 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.7E-7 | 476 | 524 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0007165 | Biological Process | signal transduction | ||||
| GO:0009658 | Biological Process | chloroplast organization | ||||
| GO:0009910 | Biological Process | negative regulation of flower development | ||||
| GO:0010380 | Biological Process | regulation of chlorophyll biosynthetic process | ||||
| GO:0010638 | Biological Process | positive regulation of organelle organization | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:1900056 | Biological Process | negative regulation of leaf senescence | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 763 aa Download sequence Send to blast |
MVIGMLAVSS LRDTYRDEKQ GGETVETSNI NFSINAGDGF PDFADCIDFD DLFIGINVDG 60 DVLPDLEMDS EILADFSAED YNSDVNASPS SAEKSADIDD NSVSINSTIT SSTTTHRAEE 120 EDDKKIKFTT TENEDDKVSG SGSGLDSSLS GSVTGRNVAS TNTNKEAADK GGGKKSSSST 180 AQSKNNNSQG KRKVKVDWTP ELHRRFVQAV EQLGVDKAVP SRILELMGID CLTRHNIASH 240 LQKYRSHRKH LLAREAEAAS WSQRRQMFGP AGGGGGGAVS GKRDVGPWPV LPTMGFPPMT 300 PPPVPALVPP PMHHHFRPLH VWGHPTVDQS LMHVWPPKHL PHSPSHHPPP QAWPPVPPPP 360 PPQDPSYWHS HHQRVPSTLT PGTPCFPPPM ATPRFGTPTV PGIPPPHAMY KVEPGIGNLN 420 PGQSGPHPLF DFHPSKESVD AAIGDVLAKP WLPLPLGLKP PSTDSVMGEL QRQGVDWTPE 480 LHRRFVQAVE QLGVDKAVPS RILELMGIDC LTRHNIASHL QKYRSHRKHL LAREAEAASW 540 SQRRQMFGPA GGGGGGAVSG KRDVGPWPVL PTMGFPPMTP PPVPPLVPPP MHHHFRPLHV 600 WGHPTVDQSL MHVWPPKHLP HSPSHHPPPQ AWPPVPPPPP PQDPSYWHSH HQRVPSTLTP 660 GTPCFPQPMA TPRFATPTVP GIPPPHAMYK VEPGIGNLNP GQSGPHPLFD FHPSKESVDA 720 AIGDVLAKPW LPLPLGLKPP STDSVMGELQ RQGVPKIPPS CA* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 6e-16 | 190 | 524 | 2 | 57 | ARR10-B |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that functions with GLK2 to promote chloroplast development. Acts as an activator of nuclear photosynthetic genes involved in chlorophyll biosynthesis, light harvesting, and electron transport. Acts in a cell-autonomous manner to coordinate and maintain the photosynthetic apparatus within individual cells. May function in photosynthetic capacity optimization by integrating responses to variable environmental and endogenous cues (PubMed:11828027, PubMed:12220263, PubMed:17533111, PubMed:18643989, PubMed:19376934, PubMed:19383092, PubMed:19726569). Prevents premature senescence (PubMed:23459204). {ECO:0000269|PubMed:11828027, ECO:0000269|PubMed:12220263, ECO:0000269|PubMed:17533111, ECO:0000269|PubMed:18643989, ECO:0000269|PubMed:19376934, ECO:0000269|PubMed:19383092, ECO:0000269|PubMed:19726569, ECO:0000269|PubMed:23459204}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00022 | PBM | Transfer from AT2G20570 | Download |
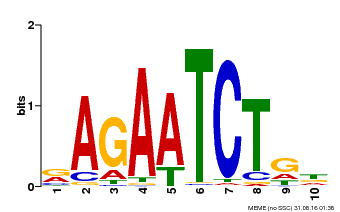
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By light. Repressed by BZR2. {ECO:0000269|PubMed:12220263, ECO:0000269|PubMed:21214652}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010093955.1 | 1e-155 | transcription activator GLK1 isoform X2 | ||||
| Swissprot | Q9SIV3 | 1e-86 | GLK1_ARATH; Transcription activator GLK1 | ||||
| TrEMBL | A0A2P5B6N0 | 0.0 | A0A2P5B6N0_PARAD; Octamer-binding transcription factor | ||||
| STRING | XP_010093955.1 | 1e-154 | (Morus notabilis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF4242 | 34 | 61 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G20570.1 | 3e-69 | GBF's pro-rich region-interacting factor 1 | ||||



