 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | KHN48291.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Glycine; Soja
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 699aa MW: 77572.3 Da PI: 7.8349 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 73.1 | 3.5e-23 | 117 | 218 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT.......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSEE.. CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrsefel 92
f k+lt+sd+++ g +++p+ +ae++ ++ + + +d +g++W++++iyr++++r++lt+GW+ Fv+ ++L +gD++vF + +++ l
KHN48291.1 117 FAKTLTQSDANNGGGFSVPRYCAETIfprldysADPPVQ--NILAKDVHGETWKFRHIYRGTPRRHLLTTGWSTFVNHKKLVAGDSIVFL--RAENGDL 211
89****************************997656666..69999********************************************..4488999 PP
EEEEE-S CS
B3 93 vvkvfrk 99
+v+++r+
KHN48291.1 212 CVGIRRA 218
****997 PP
| |||||||
| 2 | Auxin_resp | 110.3 | 1.9e-36 | 286 | 369 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrl.sGtvvgvsdldpvrWpnSkWrsLk 83
aa+ a++k++FevvY+Prast+eF+vk++ ve+a+++++ G Rfkmafetedss++++ +Gt+++v+ +dp++WpnS+Wr+L+
KHN48291.1 286 AANLAANKKPFEVVYYPRASTPEFCVKASLVEAAMQTRWYSGIRFKMAFETEDSSRISWfMGTISSVQVADPLNWPNSPWRLLQ 369
7899******************************************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101936 | 6.54E-29 | 109 | 218 | IPR015300 | DNA-binding pseudobarrel domain |
| Gene3D | G3DSA:2.40.330.10 | 1.3E-36 | 110 | 220 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 5.64E-21 | 116 | 217 | No hit | No description |
| PROSITE profile | PS50863 | 13.014 | 117 | 219 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 6.0E-21 | 117 | 219 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 2.1E-21 | 117 | 218 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 5.5E-32 | 286 | 369 | IPR010525 | Auxin response factor |
| PROSITE profile | PS51745 | 18.079 | 613 | 693 | IPR000270 | PB1 domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007389 | Biological Process | pattern specification process | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0048829 | Biological Process | root cap development | ||||
| GO:0051301 | Biological Process | cell division | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 699 aa Download sequence Send to blast |
MITFMDTKEK LKEVERCLDP QLWHACAGGM VQMPTVNTKV YYFPQGHAEH ACGPVNFKTC 60 PKVPPFVPCR VVAVKYMADP ETDEVYAKLK LVPLNANDVD YDHDVIGAET RDKPASFAKT 120 LTQSDANNGG GFSVPRYCAE TIFPRLDYSA DPPVQNILAK DVHGETWKFR HIYRGTPRRH 180 LLTTGWSTFV NHKKLVAGDS IVFLRAENGD LCVGIRRAKK GIGGGLETSS GWNPAGGNFP 240 MPYSGFSPFL REDDNRILRN GNSNGLNPSV SMMGKGKVRP EAIIEAANLA ANKKPFEVVY 300 YPRASTPEFC VKASLVEAAM QTRWYSGIRF KMAFETEDSS RISWFMGTIS SVQVADPLNW 360 PNSPWRLLQV TWDEPDLLQN VRRVSPWLVE LVSNMPAIHF SPFSPPRKKL RLPQHPDFPL 420 DGQIPLPTLP NNLLGPNNTN QFGCLLESTP AGMQGARHAH YGLSLSDLHL SKLQSGLSSA 480 GFPPLDHAAT PMKVSNNKTL QKPSISENVS CLLSMTNSTR SSKKLDAGKT PSLMLFGQKI 540 LTEQQISPSS SVDTLSPVLT RNCSTDGNVN KVTNFFDGFG SALHQQGLHE HSSCERFQWC 600 KDNHQEIEAN METGHCKVFM ESEDVGRTMD LSLLRSYDEL HRKLADMFGI EKSEMLSRVL 660 YCDSVGAIKH IGDEPFSDFT RTAKRLTILM DSGSNNVGV |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldu_A | 5e-77 | 18 | 390 | 51 | 388 | Auxin response factor 5 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00461 | DAP | Transfer from AT4G30080 | Download |
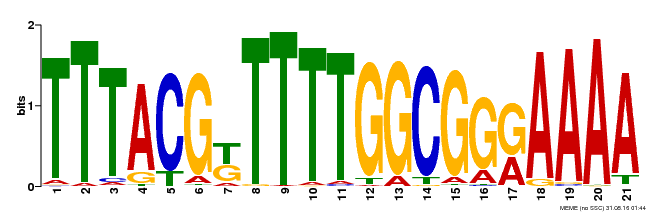
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | KHN48291.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_028198002.1 | 0.0 | auxin response factor 18-like | ||||
| Refseq | XP_028198003.1 | 0.0 | auxin response factor 18-like | ||||
| Swissprot | Q653H7 | 0.0 | ARFR_ORYSJ; Auxin response factor 18 | ||||
| TrEMBL | A0A445HFS9 | 0.0 | A0A445HFS9_GLYSO; Auxin response factor | ||||
| STRING | GLYMA10G06080.2 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF637 | 34 | 138 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G30080.1 | 0.0 | auxin response factor 16 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



