 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | KHN39228.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Glycine; Soja
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1079aa MW: 121228 Da PI: 6.0457 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 181 | 1.4e-56 | 21 | 136 | 3 | 118 |
CG-1 3 kekkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqrrcywl 101
+ ++rwl++ ei++iL n++ +++t+e+++rp+sgsl+L++rk++ryfrkDG++w+kkkdgktv+E+hekLKvg+v+vl+cyYah+een++fqrr+yw+
KHN39228.1 21 EAQHRWLRPAEICEILRNYRMFQITSEPPNRPPSGSLFLFDRKVLRYFRKDGHNWRKKKDGKTVKEAHEKLKVGSVDVLHCYYAHGEENENFQRRSYWM 119
349************************************************************************************************ PP
CG-1 102 Leeelekivlvhylevk 118
Le ++ +iv+vhyl+vk
KHN39228.1 120 LELDMMHIVFVHYLDVK 136
**************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 82.157 | 15 | 141 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 4.0E-82 | 18 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 1.4E-50 | 21 | 135 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 4.1E-5 | 493 | 581 | IPR013783 | Immunoglobulin-like fold |
| Pfam | PF01833 | 7.0E-6 | 495 | 580 | IPR002909 | IPT domain |
| SuperFamily | SSF81296 | 1.17E-16 | 495 | 581 | IPR014756 | Immunoglobulin E-set |
| SuperFamily | SSF48403 | 1.38E-15 | 685 | 789 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 1.2E-15 | 689 | 790 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 3.08E-12 | 693 | 783 | No hit | No description |
| PROSITE profile | PS50297 | 16.573 | 695 | 803 | IPR020683 | Ankyrin repeat-containing domain |
| SMART | SM00015 | 4.2 | 902 | 924 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.206 | 903 | 932 | IPR000048 | IQ motif, EF-hand binding site |
| SuperFamily | SSF52540 | 7.43E-7 | 904 | 953 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| Pfam | PF00612 | 0.0075 | 905 | 923 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0071 | 925 | 947 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.285 | 926 | 950 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 2.3E-4 | 928 | 947 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0016874 | Molecular Function | ligase activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1079 aa Download sequence Send to blast |
MSERSSFGLG PRLDLQQLQL EAQHRWLRPA EICEILRNYR MFQITSEPPN RPPSGSLFLF 60 DRKVLRYFRK DGHNWRKKKD GKTVKEAHEK LKVGSVDVLH CYYAHGEENE NFQRRSYWML 120 ELDMMHIVFV HYLDVKVNKT NIGGKTYSDE VTSDSQKSSS LSSGFPRNYG SVPSGSTDSM 180 SPTSTLTSLC EDADSEDIHQ ASSGLHSYRE SQNLGNDRPM DKIHARSNSS YLMHPFSDNH 240 GQLPVSGAEY IPHVQGNKSR ASDTTYIEGQ RAHGIASWDN AMEQSAGKHA DPSLVSSTSI 300 PSSAMGNILD KNHTVPGNLL GHKIALTEVE RGAQPVQSNW QIPFEDNTGE LPNWGFTQSL 360 GLEFGSDYGT SLLGDVTNNA GPEIDPELFT FNGELKEQYT HGQSQPALKS NSAYEVPGEA 420 SINYALTMRR GLLDGEESLK KVDSFSRWMT KELAGVDDLH MQSSPGISWS TDECGDVIDD 480 TSLHLSLSQD QLFSINDFSP KWAYAESEIE VLIVGTFLKS QPVVAKCNWS CMFGEVEVPA 540 EVLADGILCC QAPPHKIGRV PFYVTCSNRF ACSEVREFEY REGFDRNINF PDFFNNSSEM 600 ELHLRLVGLL SLNSMHTLNQ VFEGDMDKRN LIFKLISLKE EEEYSSKEET TAEMDISQQK 660 LKEHMFHKQV KEKLYSWLLH KVTETGKGPL VLDEEGQGVL HLIAALGYDW AINPIITAGV 720 NINFRDVNGW TALHWAAFCG RERTVAVLVS MDAAAGALTD PCPEFPLGRT PADLASSKGH 780 KGISGFLAES LLTSHLESLT MDENKDGRKE TSGMKVVQTV SERTATPVLN GDIPDDICLK 840 DSLNAVRNAT QAADRIYQVF RMQSFQRKQL ALYEDDEFGL SDQQALSLLA SKACRSGQGE 900 GLANAAAIQI QKKFRGWTKR KEFLIIRQRI VKIQAHVRGH QVRKQYKPII WSVGILEKVI 960 LRWRRKGSGL RGFRPASQNK VPEQPSESPK QDDYDYLKEG RKQSEVKFKK ALSRVKSMVQ 1020 YPEARAQYRR VLNVVEDFRQ TKGGNLNLIN SEETVDGVED LIDIDMLLDD ENFLPIAFD |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3' (By similarity). Regulates transcriptional activity in response to calcium signals (Probable). Binds calmodulin in a calcium-dependent manner (By similarity). Involved in freezing tolerance in association with CAMTA1 and CAMTA3. Contributes together with CAMTA1 and CAMTA3 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:23581962). Involved together with CAMTA3 and CAMTA4 in the positive regulation of a general stress response (PubMed:25039701). Involved in tolerance to aluminum. Binds to the promoter of ALMT1 transporter and contributes to the positive regulation of aluminum-induced expression of ALMT1 (PubMed:25627216). {ECO:0000250|UniProtKB:Q8GSA7, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25627216, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
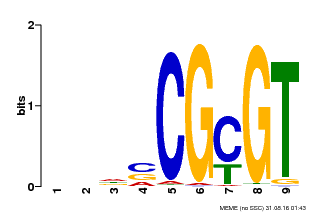
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | KHN39228.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salt, wounding, abscisic acid, H(2)O(2) and salicylic acid (PubMed:12218065). Induced by aluminum (PubMed:25627216). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:25627216}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP015034 | 0.0 | AP015034.1 Vigna angularis var. angularis DNA, chromosome 1, almost complete sequence, cultivar: Shumari. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_028243095.1 | 0.0 | calmodulin-binding transcription activator 2-like isoform X1 | ||||
| Swissprot | Q6NPP4 | 0.0 | CMTA2_ARATH; Calmodulin-binding transcription activator 2 | ||||
| TrEMBL | A0A0B2S3K7 | 0.0 | A0A0B2S3K7_GLYSO; Calmodulin-binding transcription activator 2 | ||||
| STRING | GLYMA08G07680.2 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF7406 | 26 | 42 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G64220.2 | 0.0 | Calmodulin-binding transcription activator protein with CG-1 and Ankyrin domains | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



