 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | KHN34531.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Glycine; Soja
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 205aa MW: 23346.6 Da PI: 9.8802 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 56.7 | 5.8e-18 | 85 | 135 | 2 | 55 |
AP2 2 gykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
+y+GVr++ +g+++AeIrdp++ng +r++lg++ t+e A a+++a+ k++g
KHN34531.1 85 HYRGVRRRT-WGKFAAEIRDPKKNG--ARIWLGTYETEEAAGLAYDRAAFKMRG 135
8*****877.***********9998..**********99999**********98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:3.30.730.10 | 7.6E-31 | 84 | 144 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 7.14E-29 | 84 | 145 | No hit | No description |
| SMART | SM00380 | 3.0E-35 | 85 | 149 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 23.354 | 85 | 143 | IPR001471 | AP2/ERF domain |
| Pfam | PF00847 | 1.6E-12 | 85 | 135 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 2.62E-22 | 85 | 144 | IPR016177 | DNA-binding domain |
| PRINTS | PR00367 | 4.2E-13 | 86 | 97 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 4.2E-13 | 125 | 145 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0001944 | Biological Process | vasculature development | ||||
| GO:0009864 | Biological Process | induced systemic resistance, jasmonic acid mediated signaling pathway | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0051301 | Biological Process | cell division | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 205 aa Download sequence Send to blast |
MATTISCDFS SMESIQHYLL EPEHDSNALM NISNHEVCYS ATYNAFPSPT FSLEGVVHDA 60 KDSLACKREH PREEREVHAP PVWKHYRGVR RRTWGKFAAE IRDPKKNGAR IWLGTYETEE 120 AAGLAYDRAA FKMRGSKAKL NFPHLIGSHA PPQPVRVTKA LKRNSPGKRK NLADLLNRLA 180 KNRSQILIQV LEMGLHANYV NVEQW |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2gcc_A | 5e-30 | 84 | 148 | 4 | 68 | ATERF1 |
| 3gcc_A | 5e-30 | 84 | 148 | 4 | 68 | ATERF1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Acts as a transcriptional activator. Binds to the GCC-box pathogenesis-related promoter element. Involved in the regulation of gene expression by stress factors and by components of stress signal transduction pathways. Involved in disease resistance pathways. {ECO:0000269|PubMed:10715325, ECO:0000269|PubMed:12805630, ECO:0000269|PubMed:9756931}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00548 | DAP | Transfer from AT5G47220 | Download |
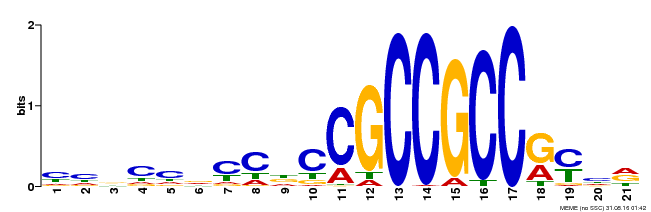
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | KHN34531.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by jasmonate (JA) and Alternaria brassicicola (locally and systemically). Ethylene induction is completely dependent on a functional ETHYLENE-INSENSITIVE2 (EIN2) while wounding induction does not require EIN2. Transcripts accumulate strongly in cycloheximide-treated plants, a protein synthesis inhibitor. Seems to not be influenced by exogenous abscisic acid (ABA), cold, heat, NaCl or drought stress. {ECO:0000269|PubMed:10715325, ECO:0000269|PubMed:12805630}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT092019 | 1e-145 | BT092019.1 Soybean clone JCVI-FLGm-9K2 unknown mRNA. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_028240846.1 | 1e-154 | ethylene-responsive transcription factor 2-like | ||||
| Swissprot | O80338 | 5e-36 | EF101_ARATH; Ethylene-responsive transcription factor 2 | ||||
| TrEMBL | A0A0B2RIU9 | 1e-153 | A0A0B2RIU9_GLYSO; Ethylene-responsive transcription factor 2 | ||||
| STRING | GLYMA07G14060.1 | 1e-152 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF512 | 33 | 157 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G47220.1 | 2e-38 | ethylene responsive element binding factor 2 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



