 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | KHN31935.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Glycine; Soja
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 267aa MW: 29943.8 Da PI: 9.1762 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 66.2 | 6.6e-21 | 118 | 168 | 2 | 55 |
AP2 2 gykGVrwdkkrgrWvAeIrdpseng.krkrfslgkfgtaeeAakaaiaarkkleg 55
+y+GVr+++ +g+++AeIrdp + k r++lg+f+ta eAaka+++a+ k++g
KHN31935.1 118 HYRGVRRRP-WGKYAAEIRDP---NrKGSRVWLGTFDTAIEAAKAYDKAAFKMRG 168
8********.**********9...6566*************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00018 | 5.48E-32 | 117 | 176 | No hit | No description |
| Gene3D | G3DSA:3.30.730.10 | 5.6E-33 | 117 | 177 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 2.68E-23 | 118 | 178 | IPR016177 | DNA-binding domain |
| SMART | SM00380 | 2.6E-38 | 118 | 182 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 24.434 | 118 | 176 | IPR001471 | AP2/ERF domain |
| Pfam | PF00847 | 1.0E-14 | 118 | 168 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 1.2E-13 | 119 | 130 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 1.2E-13 | 158 | 178 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 267 aa Download sequence Send to blast |
MQSSISQSEI CITDYLLPQE VPSQFQFPDM SNNNIPMNHT NLQMPQITSF SKPPRSSSNL 60 SNRKPSLRNI TIPSITSGLT TTMSQTITTT TTIATTMYNN NQVTSSSDET NNIKENKHYR 120 GVRRRPWGKY AAEIRDPNRK GSRVWLGTFD TAIEAAKAYD KAAFKMRGSK AILNFPLEIG 180 ESEESVSSCI KVGVKREREE ESKSNNYEKS EFNNNNNSNK HVKKEECSPK AVCPLTPSCW 240 KGFWDTDVMG TIFSVPPLSP LSPLMVV |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2gcc_A | 1e-32 | 115 | 183 | 2 | 70 | ATERF1 |
| 3gcc_A | 1e-32 | 115 | 183 | 2 | 70 | ATERF1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Acts as a transcriptional activator. Binds to the GCC-box pathogenesis-related promoter element. Involved in the regulation of gene expression by stress factors and by components of stress signal transduction pathways mediated by ethylene, that seems to depend on a protein kinase/phosphatase cascade, and to be influenced by methyl-jasmonate. {ECO:0000269|PubMed:10652129, ECO:0000269|PubMed:10792818, ECO:0000269|PubMed:7756828, ECO:0000269|Ref.2}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00556 | DAP | Transfer from AT5G51190 | Download |
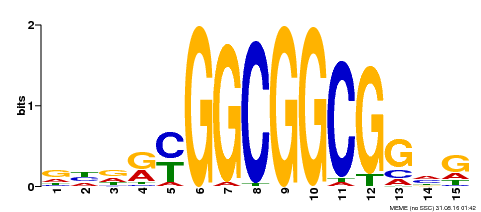
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | KHN31935.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Strongly induced by ethephon (ethylene-releasing compound) in buds and in leaves. Also strongly induced by cycloheximide, mechanical stimuli. Wounding leads to a both local and systemic expression, independently of ethylene, and through a de-novo-protein-synthesis-independent regulation. Wound induction is reduced by methyl-jasmonate. Induction by purified xylanase from Trichoderma viride (TvX) and another elicitor from Phytophthora infestans (PiE), that appears to be mediated by a protein kinase cascade, and to be negatively regulated by protein phosphatases. {ECO:0000269|PubMed:10652129, ECO:0000269|PubMed:12090623, ECO:0000269|PubMed:7756828, ECO:0000269|Ref.2}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT094806 | 0.0 | BT094806.1 Soybean clone JCVI-FLGm-6O21 unknown mRNA. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_028221868.1 | 0.0 | ethylene-responsive transcription factor ERF105-like | ||||
| Swissprot | Q40478 | 1e-54 | ERF5_TOBAC; Ethylene-responsive transcription factor 5 | ||||
| TrEMBL | A0A0B2RF45 | 0.0 | A0A0B2RF45_GLYSO; Ethylene-responsive transcription factor 5 | ||||
| STRING | GLYMA20G16910.1 | 1e-178 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1085 | 34 | 109 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G51190.1 | 7e-35 | ERF family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



