 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | KHN09491.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Glycine; Soja
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 514aa MW: 55810.4 Da PI: 9.807 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 16.1 | 3.1e-05 | 67 | 89 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
+ C++C+k F r nL+ H r H
KHN09491.1 67 FICEICNKGFQRDQNLQLHRRGH 89
89*******************88 PP
| |||||||
| 2 | zf-C2H2 | 13.9 | 0.00016 | 143 | 165 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
+kC++C+k + +s+ k H +t+
KHN09491.1 143 WKCEKCSKKYAVQSDWKAHTKTC 165
58*******************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF57667 | 4.07E-7 | 66 | 89 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.4E-5 | 66 | 89 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 10.845 | 67 | 89 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.022 | 67 | 89 | IPR015880 | Zinc finger, C2H2-like |
| Pfam | PF12171 | 6.2E-5 | 67 | 89 | IPR022755 | Zinc finger, double-stranded RNA binding |
| PROSITE pattern | PS00028 | 0 | 69 | 89 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 98 | 108 | 138 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 4.6E-5 | 131 | 164 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 4.07E-7 | 138 | 163 | No hit | No description |
| SMART | SM00355 | 130 | 143 | 163 | IPR015880 | Zinc finger, C2H2-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010075 | Biological Process | regulation of meristem growth | ||||
| GO:0045604 | Biological Process | regulation of epidermal cell differentiation | ||||
| GO:0048364 | Biological Process | root development | ||||
| GO:0051302 | Biological Process | regulation of cell division | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 514 aa Download sequence Send to blast |
MQMIPGDPFS LSTSLPAGLP NEQSTNTNPN PNPPPPNKKK RNLPGTPDPD AEVIALSPKT 60 LMATNRFICE ICNKGFQRDQ NLQLHRRGHN LPWKLRQRSN KEVRKKVYIC PEKTCVHHDA 120 ARALGDLTGI KKHYSRKHGE KKWKCEKCSK KYAVQSDWKA HTKTCGTREY KCDCGNLFSR 180 KDSFITHRAF CDALADESSR LTSVASTSLN FKNEDATMIN TQASLSTRGL ITGHGMQNVS 240 QFGPHGFRLM NMGTDQQRPN LSLWLNQGNH HINNPLDVAL SSSSSGLPEV VHMAQANINN 300 NALIGSSSVF SNFGMPASSN SSNPNLMGKK GDGGASDLAS MYSESQNKNS NSTSPMSATA 360 LLQKAAQMGS TRSTNPSIFS GSFGVMSSSS TQSTSLNSNI ANQSCDQLNQ AFQNFNATSS 420 SSATMLGSST NFSSLTHSSN GFDQFMMQNK VEPTQLKLHH PGSNSVEHNL TRDFLGVSGN 480 GGGHVHHAFL PQELAKFASL GSSMGLNQFT VGHQ |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5b3h_C | 2e-35 | 139 | 208 | 3 | 72 | Zinc finger protein JACKDAW |
| 5b3h_F | 2e-35 | 139 | 208 | 3 | 72 | Zinc finger protein JACKDAW |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 93 | 105 | KLRQRSNKEVRKK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that, together with BIB, regulates tissue boundaries and asymmetric cell division by a rapid up-regulation of 'SCARECROW' (SCR), thus controlling the nuclear localization of 'SHORT-ROOT' (SHR) and restricting its action (PubMed:17785527, PubMed:25829440). Binds DNA via its zinc fingers (PubMed:28211915, PubMed:24821766). Recognizes and binds to SCL3 promoter sequence 5'-AGACAA-3' to promotes its expression when in complex with RGA (PubMed:24821766). Confines CYCD6 expression to the cortex-endodermis initial/daughter (CEI/CEID) tissues (PubMed:25829440). Required for radial patterning and stem cell maintenance (PubMed:17785527). Counteracted by 'MAGPIE' (MGP) (PubMed:17785527). Binds to the SCR and MGP promoter sequences (PubMed:21935722). Controls position-dependent signals that regulate epidermal-cell-type patterning (PubMed:20356954). {ECO:0000269|PubMed:17785527, ECO:0000269|PubMed:20356954, ECO:0000269|PubMed:21935722, ECO:0000269|PubMed:24821766, ECO:0000269|PubMed:25829440, ECO:0000269|PubMed:28211915}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00485 | DAP | Transfer from AT5G03150 | Download |
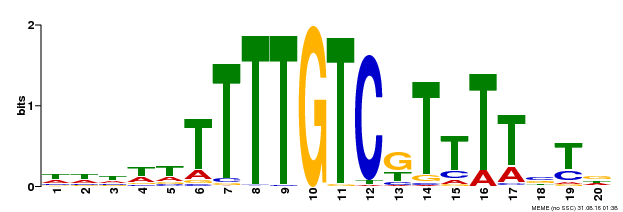
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | KHN09491.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Not regulated by SCR and SHR. {ECO:0000269|PubMed:21935722}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP015037 | 1e-165 | AP015037.1 Vigna angularis var. angularis DNA, chromosome 4, almost complete sequence, cultivar: Shumari. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_028225828.1 | 0.0 | zinc finger protein JACKDAW-like | ||||
| Refseq | XP_028225830.1 | 0.0 | zinc finger protein JACKDAW-like | ||||
| Swissprot | Q700D2 | 1e-132 | IDD10_ARATH; Zinc finger protein JACKDAW | ||||
| TrEMBL | A0A445LE53 | 0.0 | A0A445LE53_GLYSO; Zinc finger protein JACKDAW | ||||
| STRING | GLYMA03G33700.1 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF181 | 34 | 275 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G03150.1 | 1e-114 | C2H2-like zinc finger protein | ||||



