 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | KHN04419.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Glycine; Soja
|
||||||||
| Family | AP2 | ||||||||
| Protein Properties | Length: 663aa MW: 73481.8 Da PI: 7.8634 | ||||||||
| Description | AP2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 50.9 | 3.7e-16 | 297 | 356 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrd.pse.ng..kr.krfslgkfgtaeeAakaaiaarkkleg 55
s+y+GV++++++gr++A+++d ++ g ++ ++++lg ++ +e+Aa+a++ a++k++g
KHN04419.1 297 SQYRGVTRHRWTGRYEAHLWDnSCKkEGqtRKgRQVYLGGYDMEEKAARAYDLAALKYWG 356
78*******************988886678446*************************98 PP
| |||||||
| 2 | AP2 | 46.8 | 7.6e-15 | 399 | 450 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
s y+GV+++++ grW A+I + +k +lg+f t eeAa+a++ a+ k++g
KHN04419.1 399 SIYRGVTRHHQHGRWQARIGRVAG---NKDLYLGTFSTQEEAAEAYDVAAIKFRG 450
57****************988532...5*************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF00847 | 9.1E-13 | 297 | 356 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 2.68E-16 | 297 | 366 | IPR016177 | DNA-binding domain |
| CDD | cd00018 | 4.21E-21 | 297 | 363 | No hit | No description |
| Gene3D | G3DSA:3.30.730.10 | 3.1E-15 | 298 | 364 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 1.1E-27 | 298 | 370 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 18.782 | 298 | 364 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 1.5E-6 | 299 | 310 | IPR001471 | AP2/ERF domain |
| Pfam | PF00847 | 3.2E-10 | 399 | 450 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 2.79E-26 | 399 | 460 | No hit | No description |
| SuperFamily | SSF54171 | 5.49E-18 | 399 | 460 | IPR016177 | DNA-binding domain |
| Gene3D | G3DSA:3.30.730.10 | 5.3E-19 | 400 | 458 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 3.3E-34 | 400 | 464 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 19.137 | 400 | 458 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 1.5E-6 | 440 | 460 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007276 | Biological Process | gamete generation | ||||
| GO:0010492 | Biological Process | maintenance of shoot apical meristem identity | ||||
| GO:0042127 | Biological Process | regulation of cell proliferation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 663 aa Download sequence Send to blast |
MKRINESNNT DDGNNHNWLG FSLSPHMKME ATSAPTVPTT FYMSPSQSHL SNFGMCYGVG 60 ENGNFHSPLT VMPLKSDGSL CILEALKRSQ TQVMVPTSSP KLEDFLGGAT MGTHEYGSHE 120 RGLSLDSIYY NSQNAEAQPN RDLLSQPFRQ QGHMSVQTHP YYSGLACHGL YQAPLEEETT 180 KETHVSDCSS LMPQMTEGLK NWVAPTREFS THQQVLEQQM NCGMGNERNG VSLGSVGCGE 240 LQSLSLSMSP GSQSSCVTAP SGTDSVAVDA KKRGHAKLGQ KQPVHRKSID TFGQRTSQYR 300 GVTRHRWTGR YEAHLWDNSC KKEGQTRKGR QVYLGGYDME EKAARAYDLA ALKYWGPSTH 360 INFSIENYQV QLEEMKNMSR QEYVAHLRRK SSGFSRGASI YRGVTRHHQH GRWQARIGRV 420 AGNKDLYLGT FSTQEEAAEA YDVAAIKFRG ANAVTNFDIS RYDVERIMAS SNLLAGELAR 480 RKKDNDPRNK DIDYNKSVVT SVNNEETVQV QAGNNNNEND SEWKMVLFNH PSQQQQANGN 540 GSDQKIMNCG NYRNSAFSMA LQDLIGIDSV GSGQHNMLDE SSKIGTHFSN TSSLVTSLSS 600 SREASPEKRG PSLLFPMPPM ETKIVNPIGT SVTSWLPSPT VQMRPSPAIS LSHLPVFASW 660 TDT |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CAC[AG]N[AT]TNCCNANG-3'. Required for the initiation and growth of ovules integumenta, and for the development of female gametophyte. Plays a critical role in the development of gynoecium marginal tissues (e.g. stigma, style and septa), and in the fusion of carpels and of medial ridges leading to ovule primordia. Also involved in organs initiation and development, including floral organs. Maintains the meristematic competence of cells and consequently sustains expression of cell cycle regulators during organogenesis, thus controlling the final size of each organ by controlling their cell number. Regulates INO autoinduction and expression pattern. As ANT promotes petal cell identity and mediates down-regulation of AG in flower whorl 2, it functions as a class A homeotic gene. {ECO:0000269|PubMed:10528263, ECO:0000269|PubMed:10639184, ECO:0000269|PubMed:10948255, ECO:0000269|PubMed:11041883, ECO:0000269|PubMed:12183381, ECO:0000269|PubMed:12271029, ECO:0000269|PubMed:12655002, ECO:0000269|PubMed:8742706, ECO:0000269|PubMed:8742707, ECO:0000269|PubMed:9001406, ECO:0000269|PubMed:9093862, ECO:0000269|PubMed:9118807}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00088 | SELEX | Transfer from AT4G37750 | Download |
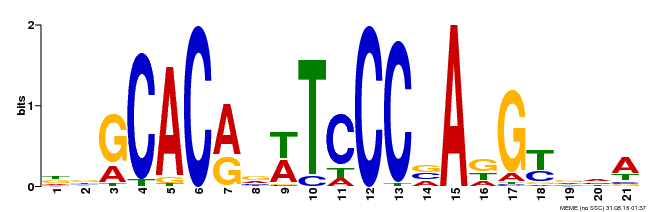
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | KHN04419.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP015035 | 1e-123 | AP015035.1 Vigna angularis var. angularis DNA, chromosome 2, almost complete sequence, cultivar: Shumari. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_028227754.1 | 0.0 | AP2-like ethylene-responsive transcription factor ANT | ||||
| Swissprot | Q38914 | 1e-159 | ANT_ARATH; AP2-like ethylene-responsive transcription factor ANT | ||||
| TrEMBL | A0A445KVX4 | 0.0 | A0A445KVX4_GLYSO; AP2-like ethylene-responsive transcription factor ANT isoform A | ||||
| STRING | GLYMA04G05080.2 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF916 | 34 | 114 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G37750.1 | 1e-152 | AP2 family protein | ||||



