 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Gorai.011G008700.6 | ||||||||
| Common Name | B456_011G008700 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Malvoideae; Gossypium
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 527aa MW: 57545.3 Da PI: 8.5969 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 35.6 | 2.2e-11 | 479 | 524 | 4 | 47 |
S-HHHHHHHHHHHHHTTTT-HHHHHHHHT...TTS-HHHHHHHHHHH CS
Myb_DNA-binding 4 WTteEdellvdavkqlGggtWktIartmg...kgRtlkqcksrwqky 47
WT eE++ l+++ +++G+ Wk+I + k+Rt ++k++w+++
Gorai.011G008700.6 479 WTHEEEQALIKGYREYGTQ-WKLILESYRdilKKRTQVDLKDKWRNL 524
*****************99.*************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 14.223 | 471 | 526 | IPR017930 | Myb domain |
| SMART | SM00717 | 0.0016 | 475 | 526 | IPR001005 | SANT/Myb domain |
| SuperFamily | SSF46689 | 2.94E-11 | 477 | 524 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 2.2E-10 | 479 | 524 | IPR009057 | Homeodomain-like |
| Pfam | PF13921 | 3.0E-9 | 479 | 525 | No hit | No description |
| CDD | cd11660 | 1.89E-16 | 479 | 526 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009901 | Biological Process | anther dehiscence | ||||
| GO:0010152 | Biological Process | pollen maturation | ||||
| GO:0043067 | Biological Process | regulation of programmed cell death | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 527 aa Download sequence Send to blast |
MLPPTVDNVV YASAQVPSSP HPCSDEGKMG NVVSASAEAS PSPHPCADKG NVDNVISSSA 60 QVPSSPHPCT NKKKKIRRAS FQARHKPRPR CKQPRGVVIT DMEEDQPLCT KNGTPSSFEV 120 NGCLQVASKT RSAALLDGVT DSPSEALEVL ESVALVMAGK NFHPKGSVED SNKDKGDPTT 180 SFYPTTHHCT EKGKQILMVN SQTSGNPNSS HRHCEGPAAT ADIEGYRTLS TPDIDKLRDA 240 LTSSIADLTA TVADPLPKAL EVAATLVSFM EAAKSLSTDG AEGDVRKEKG VPAAPMNCNF 300 EQAQAERRAP DKAIREDQDK MARSVPASSV GHSVEPSQAK EGNETFSRPK KVPKRNLMKR 360 NDAAHTHERR APYEGIREDY NKVDGSVPAP SVNHTAEPSH SKEGNEAVIR PKKVRKRSIM 420 ERNDTAHASE WEDSIDGSDG GTSGCSNRCT LPSPKTNHVS PLKESEPQKQ KNRRKTNWWT 480 HEEEQALIKG YREYGTQWKL ILESYRDILK KRTQVDLKDK WRNLSK* |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00052 | PBM | Transfer from AT1G15720 | Download |
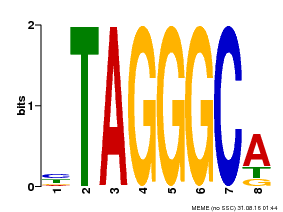
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_012454054.1 | 0.0 | PREDICTED: uncharacterized protein LOC105776112 isoform X1 | ||||
| TrEMBL | A0A0D2RF44 | 0.0 | A0A0D2RF44_GOSRA; Uncharacterized protein | ||||
| STRING | Gorai.011G008700.1 | 0.0 | (Gossypium raimondii) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G15720.1 | 2e-13 | TRF-like 5 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Gorai.011G008700.6 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



