 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Gorai.011G001500.1 | ||||||||
| Common Name | B456_011G001500, LOC105776298 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Malvoideae; Gossypium
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 434aa MW: 48098.9 Da PI: 6.0405 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 106.5 | 1.5e-33 | 227 | 281 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
kpr+rWtpeLHerFveav++L G+ekAtPk +l+lm+v+gLt++hvkSHLQkYRl
Gorai.011G001500.1 227 KPRMRWTPELHERFVEAVNKLHGPEKATPKGVLKLMNVEGLTIYHVKSHLQKYRL 281
79****************************************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 11.732 | 224 | 284 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 3.23E-18 | 225 | 281 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 5.6E-25 | 227 | 282 | IPR006447 | Myb domain, plants |
| Gene3D | G3DSA:1.10.10.60 | 3.2E-31 | 227 | 282 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 8.5E-10 | 229 | 280 | IPR001005 | SANT/Myb domain |
| Pfam | PF14379 | 1.6E-22 | 318 | 365 | IPR025756 | MYB-CC type transcription factor, LHEQLE-containing domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 434 aa Download sequence Send to blast |
MTQSEPSKGI GERYPIAAFP AHKILSIKSE GQSSMAGECS SRLSFPFNSP RSDAKNASSS 60 SSVFCTSLYL SSSSTSETQR QLGNLPFLPH PPTRHQFISS VDSSKSPAMF SEDLNNPYNE 120 GHSEIIMKDF LNLPADASSD GCFYGMHCES DDFGITTEQL ELQFLSDELD IAITDHGENP 180 RLDEIYETCP PSSAKPTVEL KCNQNTNSVS PTIDGPAMSG AAAAVLKPRM RWTPELHERF 240 VEAVNKLHGP EKATPKGVLK LMNVEGLTIY HVKSHLQKYR LAKYMPEKKE ERKTSSSSEE 300 KKPASSSNES DGKKRGGMHI TEALRMQMEV QKQLHEQLEL QRSLQLRIEE HARYLQKMLE 360 EQQKAGNALI PNSNSELPSK IASETEPCES KTESPTSLPS KHKAPNVEDC KLKSSPKRLR 420 MDEAEAEVEN PEQ* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 4e-26 | 227 | 284 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 4e-26 | 227 | 284 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 4e-26 | 227 | 284 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 4e-26 | 227 | 284 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 4e-26 | 227 | 284 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 4e-26 | 227 | 284 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 4e-26 | 227 | 284 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 4e-26 | 227 | 284 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Gra.1278 | 0.0 | flower| flowering | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 48806945 | 0.0 | ||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00354 | DAP | Transfer from AT3G13040 | Download |
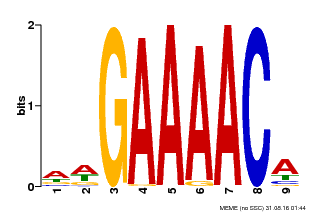
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_012454324.1 | 0.0 | PREDICTED: uncharacterized protein LOC105776298 | ||||
| Refseq | XP_012454325.1 | 0.0 | PREDICTED: uncharacterized protein LOC105776298 | ||||
| Swissprot | Q949U2 | 1e-129 | PHL6_ARATH; Myb family transcription factor PHL6 | ||||
| TrEMBL | A0A0D2VG79 | 0.0 | A0A0D2VG79_GOSRA; Uncharacterized protein | ||||
| STRING | Gorai.011G001500.1 | 0.0 | (Gossypium raimondii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM6786 | 27 | 44 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G13040.2 | 5e-95 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Gorai.011G001500.1 |
| Entrez Gene | 105776298 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



