 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Gorai.006G131700.2 | ||||||||
| Common Name | B456_006G131700, LOC105799438 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Malvoideae; Gossypium
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 506aa MW: 55592.2 Da PI: 6.3022 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 21.1 | 8.4e-07 | 263 | 284 | 2 | 23 |
EETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 2 kCpdCgksFsrksnLkrHirtH 23
C++Cgk F+r nL+ H+r H
Gorai.006G131700.2 263 FCTICGKGFKRDANLRMHMRGH 284
6*******************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF57667 | 2.85E-5 | 261 | 285 | No hit | No description |
| PROSITE profile | PS50157 | 11.946 | 262 | 289 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0026 | 262 | 284 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 1.1E-5 | 263 | 313 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 264 | 284 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 53 | 311 | 344 | IPR015880 | Zinc finger, C2H2-like |
| SMART | SM00355 | 26 | 349 | 371 | IPR015880 | Zinc finger, C2H2-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010044 | Biological Process | response to aluminum ion | ||||
| GO:0010447 | Biological Process | response to acidic pH | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 506 aa Download sequence Send to blast |
MDLKERVCGE LQKEVPKGAS FTNFNTQHQQ KWEDPSSTSD YSTRIEPPFQ AFNQTSQTQS 60 LLLCNNQIKV PMQDGSVNDL LQASKIQDWD PSAMLNNLSF LGQKIHQLQD LVYLIIGRKG 120 QVLGGPDELL VQQQQLVTAD LTSIIVQLIS TAGSLLPSVK QTLSAATTSL GQCGEFGGVV 180 FPSSAQGLND GVQPQNAGGS KVSEPPNSVD ITSNNGNEQS NHIIEEHELK DEEDAEEGEN 240 LLPGTYEILQ LEKEEILAPH THFCTICGKG FKRDANLRMH MRGHGEEYKT PGALAKPTKE 300 SSSEPTIIKR YSCPYAGCKR NKDHKKFQPL KTILCVKNHY KRTHCDKSYI CSRCNTKKFS 360 VIADLKTHEK HCGKDKWLCS CGTTFSRKDK LFGHITLFQG HTPAIPSDEI KAAPAGTSDG 420 ATNKVGSVNF NLSSYVSSES EVQSSSMDIK GGIDDAVGYF SPLSFDTCNF GGFHEFPRPP 480 FDDSESSFAF LLSRSCNYSQ KSGEE* |
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots (e.g. root tips and lateral roots), leaves, flowers (e.g. stigma, sepal, anther, and filament), stems, siliques and cotyledons. {ECO:0000269|PubMed:23935008}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. Together with STOP2, plays a critical role in tolerance to major stress factors in acid soils such as proton H(+) and aluminum ion Al(3+). Required for the expression of genes in response to acidic stress (e.g. ALMT1 and MATE), and Al-activated citrate exudation. {ECO:0000269|PubMed:17535918, ECO:0000269|PubMed:18826429, ECO:0000269|PubMed:19321711, ECO:0000269|PubMed:23935008}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00196 | ampDAP | Transfer from AT1G34370 | Download |
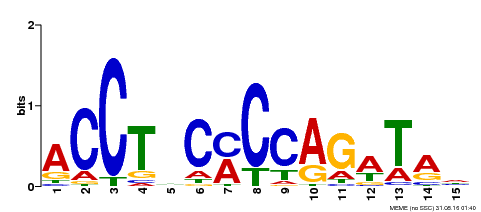
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By shock H(+) and Al(3+) treatments. {ECO:0000269|PubMed:17535918}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_012485445.1 | 0.0 | PREDICTED: protein SENSITIVE TO PROTON RHIZOTOXICITY 1-like isoform X1 | ||||
| Refseq | XP_012485446.1 | 0.0 | PREDICTED: protein SENSITIVE TO PROTON RHIZOTOXICITY 1-like isoform X1 | ||||
| Refseq | XP_012485447.1 | 0.0 | PREDICTED: protein SENSITIVE TO PROTON RHIZOTOXICITY 1-like isoform X1 | ||||
| Refseq | XP_012485448.1 | 0.0 | PREDICTED: protein SENSITIVE TO PROTON RHIZOTOXICITY 1-like isoform X1 | ||||
| Swissprot | Q9C8N5 | 0.0 | STOP1_ARATH; Protein SENSITIVE TO PROTON RHIZOTOXICITY 1 | ||||
| TrEMBL | A0A0D2RVJ5 | 0.0 | A0A0D2RVJ5_GOSRA; Uncharacterized protein | ||||
| STRING | Gorai.006G131700.1 | 0.0 | (Gossypium raimondii) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G34370.2 | 0.0 | C2H2 family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Gorai.006G131700.2 |
| Entrez Gene | 105799438 |



