 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | KXZ54938.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Chlorophyta; Chlorophyceae; Chlamydomonadales; Volvocaceae; Gonium
|
||||||||
| Family | E2F/DP | ||||||||
| Protein Properties | Length: 169aa MW: 18725.2 Da PI: 4.9257 | ||||||||
| Description | E2F/DP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | E2F_TDP | 78.3 | 7.1e-25 | 1 | 59 | 8 | 71 |
E2F_TDP 8 lltqkflkllekseegivtlnevakeLvsedvknkrRRiYDilNVLealnliekkekneirwkg 71
+lt+kfl+l+ ++e+g+++ln++a++L +v ++RRiYDi+NVLe+++liekk+kn+irwkg
KXZ54938.1 1 MLTKKFLTLIDNAEDGVLDLNKAAETL---KV--QKRRIYDITNVLEGVGLIEKKSKNNIRWKG 59
59*************************...88..****************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF02319 | 1.7E-22 | 1 | 59 | IPR003316 | E2F/DP family, winged-helix DNA-binding domain |
| SMART | SM01372 | 3.4E-26 | 1 | 59 | IPR003316 | E2F/DP family, winged-helix DNA-binding domain |
| SuperFamily | SSF46785 | 6.46E-16 | 2 | 60 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| Gene3D | G3DSA:1.10.10.10 | 4.6E-25 | 2 | 60 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| CDD | cd14660 | 2.39E-31 | 71 | 149 | No hit | No description |
| SuperFamily | SSF144074 | 4.45E-22 | 71 | 149 | No hit | No description |
| Pfam | PF16421 | 6.9E-20 | 75 | 149 | IPR032198 | E2F transcription factor, CC-MB domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008284 | Biological Process | positive regulation of cell proliferation | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0010090 | Biological Process | trichome morphogenesis | ||||
| GO:0042023 | Biological Process | DNA endoreduplication | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0051446 | Biological Process | positive regulation of meiotic cell cycle | ||||
| GO:0051782 | Biological Process | negative regulation of cell division | ||||
| GO:0005667 | Cellular Component | transcription factor complex | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0005737 | Cellular Component | cytoplasm | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| GO:0046982 | Molecular Function | protein heterodimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 169 aa Download sequence Send to blast |
MLTKKFLTLI DNAEDGVLDL NKAAETLKVQ KRRIYDITNV LEGVGLIEKK SKNNIRWKGA 60 GGGDGRESEP EVSRLRMDIR SLEDQERSLD EHIRIMSGVI QSITDNPINK NRLYVTDEDV 120 TALPAFASDT IFAVKAPPGT TLEVPDPREA ADPRDGGLRY SPAFLNAFA |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1cf7_A | 8e-24 | 1 | 61 | 14 | 75 | PROTEIN (TRANSCRIPTION FACTOR E2F-4) |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 26 | 32 | LKVQKRR |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds DNA cooperatively with DP proteins through the E2 recognition site, 5'-TTTC[CG]CGC-3' found in the promoter region of a number of genes whose products are involved in cell cycle regulation or in DNA replication. The binding of retinoblastoma-related proteins represses transactivation. Regulates gene expression both positively and negatively. Activates the expression of E2FB. Involved in the control of cell-cycle progression from G1 to S phase. Stimulates cell proliferation and delays differentiation. {ECO:0000269|PubMed:11669580, ECO:0000269|PubMed:11786543, ECO:0000269|PubMed:11862494, ECO:0000269|PubMed:11889041, ECO:0000269|PubMed:11891240, ECO:0000269|PubMed:12913157, ECO:0000269|PubMed:15377755, ECO:0000269|PubMed:16514015, ECO:0000269|PubMed:19662336}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00190 | DAP | Transfer from AT1G47870 | Download |
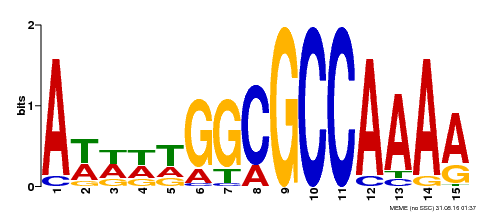
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002948442.1 | 2e-92 | E2F transcription factor family | ||||
| Swissprot | Q9FNY0 | 1e-46 | E2FA_ARATH; Transcription factor E2FA | ||||
| TrEMBL | A0A150GZ33 | 1e-119 | A0A150GZ33_GONPE; E2F1 protein | ||||
| STRING | XP_002948442.1 | 8e-92 | (Volvox carteri) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Chlorophytae | OGCP3258 | 13 | 14 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G36010.1 | 8e-50 | E2F transcription factor 3 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



