 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Glyma.18G268700.1.p | ||||||||
| Common Name | GLYMA_18G268700, LOC100816424 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Glycine; Soja
|
||||||||
| Family | TCP | ||||||||
| Protein Properties | Length: 385aa MW: 43005.4 Da PI: 9.9767 | ||||||||
| Description | TCP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | TCP | 97.7 | 2.2e-30 | 81 | 202 | 2 | 113 |
TCP 2 agkkdrhskihTkvggRdRRvRlsaecaarfFdLqdeLGfdkdsktieWLlqqakpaikeltgt..............ssssaseceaes 77
g+kdrhsk+ T +g+RdRRvRls+++a++++dLqd+LG+ ++sk ++WLl +ak++i+el+ + s ++++++
Glyma.18G268700.1.p 81 FGGKDRHSKVFTIRGLRDRRVRLSVPTAIQLYDLQDRLGLSQPSKVVDWLLDAAKHEIDELPPLpvvpsvnnftlghpS-----AVTSNE 165
689*************************************************************666555555442221.....222222 PP
TCP 78 ssssasnsssg............kaaksaakskksqksaasalnlake 113
s+s+++++ + +++++ +l++++
Glyma.18G268700.1.p 166 ATTSNSQPNEQllninrsiqwegS-----------NQNSTWKLKPKEV 202
222222222223333333334321...........2222222222222 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51369 | 29.858 | 83 | 141 | IPR017887 | Transcription factor TCP subgroup |
| Pfam | PF03634 | 1.1E-28 | 83 | 213 | IPR005333 | Transcription factor, TCP |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0030154 | Biological Process | cell differentiation | ||||
| GO:0031347 | Biological Process | regulation of defense response | ||||
| GO:0045962 | Biological Process | positive regulation of development, heterochronic | ||||
| GO:0009507 | Cellular Component | chloroplast | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 385 aa Download sequence Send to blast |
MVPWNRILTI TKWKDKTQHL ATNNERHKIM IKSPKEADFP LKQEGHSQSI TDPEKAKASA 60 SSVAQWPRLK DPRIVRVSRA FGGKDRHSKV FTIRGLRDRR VRLSVPTAIQ LYDLQDRLGL 120 SQPSKVVDWL LDAAKHEIDE LPPLPVVPSV NNFTLGHPSA VTSNEATTSN SQPNEQLLNI 180 NRSIQWEGSN QNSTWKLKPK EVSREMISTD KPNWISRSEE DKQGSNNEGP STRVIPNINL 240 LPRANLNHPS FLGLLNTMPH GYQWEPSAGD VSHQLGNNGF ANQTDMHSIN VVPFPTSTLS 300 LSTGNSQILL CPPGTAQPYF PSHVTMDMDP RQINHYQMLS SGSQNPLANY LNHSFSLAMM 360 THKPLHSPNS SKSPSHKDQD FPSN* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5zkt_A | 2e-18 | 88 | 142 | 1 | 55 | Putative transcription factor PCF6 |
| 5zkt_B | 2e-18 | 88 | 142 | 1 | 55 | Putative transcription factor PCF6 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Gma.50350 | 0.0 | flower| leaf| meristem | ||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00326 | DAP | Transfer from AT3G02150 | Download |
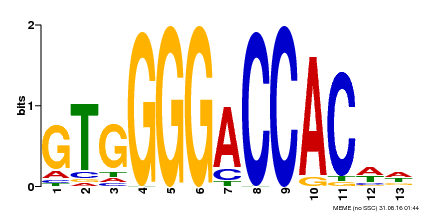
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Glyma.18G268700.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP015037 | 0.0 | AP015037.1 Vigna angularis var. angularis DNA, chromosome 4, almost complete sequence, cultivar: Shumari. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003552583.1 | 0.0 | transcription factor TCP13 | ||||
| TrEMBL | K7MUZ0 | 0.0 | K7MUZ0_SOYBN; Uncharacterized protein | ||||
| STRING | GLYMA18G50371.1 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF932 | 34 | 119 | Representative plant | OGRP180 | 15 | 163 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G02150.2 | 2e-40 | plastid transcription factor 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Glyma.18G268700.1.p |
| Entrez Gene | 100816424 |



