 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Glyma.13G187500.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Glycine; Soja
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 534aa MW: 58873.7 Da PI: 5.4557 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 56 | 9.2e-18 | 34 | 81 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT Ed++lvd+vk++G g+W+++ ++ g+ R++k+c++rw ++l
Glyma.13G187500.1.p 34 KGPWTSTEDDILVDYVKKHGEGNWNAVQKHTGLFRCGKSCRLRWANHL 81
79******************************************9986 PP
| |||||||
| 2 | Myb_DNA-binding | 45.7 | 1.5e-14 | 87 | 130 | 1 | 46 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqk 46
+g++ +eE+ l++++++++G++ W++ a++++ gRt++++k++w++
Glyma.13G187500.1.p 87 KGAFAAEEERLIAELHAKMGNK-WARMAAHLP-GRTDNEIKNYWNT 130
68899*****************.*********.***********96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 16.025 | 29 | 81 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.6E-22 | 32 | 84 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 5.8E-15 | 33 | 83 | IPR001005 | SANT/Myb domain |
| SuperFamily | SSF46689 | 3.71E-29 | 33 | 128 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 7.8E-16 | 34 | 81 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.19E-11 | 36 | 81 | No hit | No description |
| PROSITE profile | PS51294 | 23.782 | 82 | 136 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.4E-23 | 85 | 135 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 1.4E-14 | 86 | 134 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.3E-13 | 87 | 130 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 2.22E-10 | 92 | 130 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009789 | Biological Process | positive regulation of abscisic acid-activated signaling pathway | ||||
| GO:0043068 | Biological Process | positive regulation of programmed cell death | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0045926 | Biological Process | negative regulation of growth | ||||
| GO:0048235 | Biological Process | pollen sperm cell differentiation | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 534 aa Download sequence Send to blast |
MKKDIEDEVL PNDMSGAQLN DESYEGSAGI VLKKGPWTST EDDILVDYVK KHGEGNWNAV 60 QKHTGLFRCG KSCRLRWANH LRPNLKKGAF AAEEERLIAE LHAKMGNKWA RMAAHLPGRT 120 DNEIKNYWNT RIKRRQRAGL PLYPPEVSLH ALQESQHSQS TGGLNGGDKM HPDFLQKNSY 180 EIHDAIFDSL KDNQGILPYV HELSDISVYG NKLKGLDSSQ YCSFVPPTSP KRKRLKESTI 240 PFIDSCAMKK KDLYPFDQIQ DNNSDKIAQS FGMQAPLDPG LSSHSSMCYS HSLSNGNSST 300 SKPYEAMKLE LPSLQYPELD LGSWGSSPPP SLLESVDDFL QSPTAISTLE SDCSSPQNSG 360 LLDALLYQAK TMSSSKNHCS DKSSNSSTAT PGDRADSSIL NVYETEWEDY ADPVSPFGAT 420 SILNECPALR ANANSLNGQL PVQTFTGNLP KLESVDQVWT PNNENLTLSL LNITRPDFLL 480 ASDWYDLGSG HCKNQTITTD AATAFLGDDL ATDLKHMTAG ISKTMFENPF QPL* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 4e-31 | 32 | 135 | 5 | 107 | B-MYB |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Gma.24161 | 0.0 | cotyledon | ||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00490 | DAP | Transfer from AT5G06100 | Download |
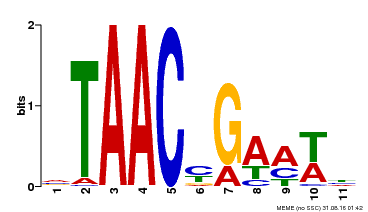
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Glyma.13G187500.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KC525897 | 0.0 | KC525897.1 Glycine max GAMBY1-like family protein (GAMYB1) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006594349.2 | 0.0 | transcription factor GAMYB-like isoform X1 | ||||
| Refseq | XP_006594350.2 | 0.0 | transcription factor GAMYB-like isoform X1 | ||||
| Refseq | XP_006594351.2 | 0.0 | transcription factor GAMYB-like isoform X1 | ||||
| Refseq | XP_006594352.2 | 0.0 | transcription factor GAMYB-like isoform X1 | ||||
| TrEMBL | A0A0R0GQI4 | 0.0 | A0A0R0GQI4_SOYBN; Uncharacterized protein | ||||
| STRING | GLYMA13G25716.2 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF2047 | 34 | 85 | Representative plant | OGRP5 | 17 | 1784 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G06100.3 | 1e-110 | myb domain protein 33 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Glyma.13G187500.1.p |



