 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Glyma.11G242200.3.p | ||||||||
| Common Name | GLYMA_11G242200 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Glycine; Soja
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 310aa MW: 35310.2 Da PI: 4.5074 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 59.5 | 5.4e-19 | 55 | 108 | 3 | 56 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k+++++ eq+++Le+ Fe ++++ e++ +LA++lgL+ rqV vWFqNrRa++k
Glyma.11G242200.3.p 55 KKRRLSVEQVKALEKNFEVENKLEPERKVKLAQELGLQPRQVAVWFQNRRARWK 108
56689************************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 130.5 | 6.5e-42 | 54 | 146 | 1 | 93 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLree 90
ekkrrls eqvk+LE++Fe e+kLeperKv+la+eLglqprqvavWFqnrRAR+ktkqlE+dy +Lk++ydalk + +L++++e+Lr++
Glyma.11G242200.3.p 54 EKKRRLSVEQVKALEKNFEVENKLEPERKVKLAQELGLQPRQVAVWFQNRRARWKTKQLERDYGVLKANYDALKLNFGTLNQDNEALRKQ 143
69**************************************************************************************** PP
HD-ZIP_I/II 91 lke 93
+ke
Glyma.11G242200.3.p 144 IKE 146
997 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 3.21E-20 | 35 | 112 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.929 | 50 | 110 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 6.1E-18 | 53 | 114 | IPR001356 | Homeobox domain |
| Pfam | PF00046 | 3.5E-16 | 55 | 108 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 6.65E-17 | 55 | 111 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 3.9E-20 | 57 | 117 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 8.6E-6 | 81 | 90 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 85 | 108 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 8.6E-6 | 90 | 106 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 8.2E-17 | 110 | 151 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009788 | Biological Process | negative regulation of abscisic acid-activated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 310 aa Download sequence Send to blast |
MLVILIMMMS DLICIGLEEH SPRNSHHMYG REFRSMLDGL DEEGCVEEPG HQSEKKRRLS 60 VEQVKALEKN FEVENKLEPE RKVKLAQELG LQPRQVAVWF QNRRARWKTK QLERDYGVLK 120 ANYDALKLNF GTLNQDNEAL RKQIKELKSR LLQEENTAGS GVSVKEEEIT TMPADSEEKT 180 MEQSKSDPPS ETSNINPSSE SSEEDHLNYE CFNNNSDDCV VGGSAAASLL QVDFMKDGSS 240 DSDGSSAILN EDTMYLPSSM NCFQFQKPYH HAQYVKTEEH NFLSADEACN FFSDEQAPTL 300 QWYCPEQWS* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 102 | 110 | RRARWKTKQ |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Gma.3154 | 0.0 | cotyledon| epicotyl| flower| hypocotyl| leaf| meristem| pod| root| stem | ||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00271 | DAP | Transfer from AT2G22430 | Download |
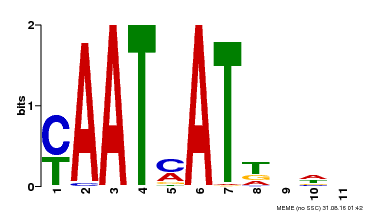
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Glyma.11G242200.3.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC235215 | 0.0 | AC235215.1 Glycine max strain Williams 82 clone GM_WBb0020M12, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003538603.1 | 0.0 | homeobox-leucine zipper protein ATHB-6 | ||||
| Refseq | XP_028196075.1 | 0.0 | homeobox-leucine zipper protein ATHB-6-like | ||||
| Refseq | XP_028196076.1 | 0.0 | homeobox-leucine zipper protein ATHB-6-like | ||||
| TrEMBL | I1LNH9 | 0.0 | I1LNH9_SOYBN; Uncharacterized protein | ||||
| STRING | GLYMA11G37920.5 | 0.0 | (Glycine max) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22430.1 | 5e-66 | homeobox protein 6 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Glyma.11G242200.3.p |



